

| Basic Information | |
|---|---|
| Species | Carica papaya |
| Cazyme ID | evm.model.supercontig_2.357 |
| Family | AA1 |
| Protein Properties | Length: 432 Molecular Weight: 48565.7 Isoelectric Point: 8.8528 |
| Chromosome | Chromosome/Scaffold: 2 Start: 4086397 End: 4088225 |
| Description | laccase 13 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA1 | 32 | 363 | 0 |
| AETHYHEFVIEERPVKRLCKTHGRITVNGQFPGPTLEVRDGDSLVITAINKAQYNVTLHWHGIRQLRNPWADGPEYITQCPILPGRSYTYRFTIINQEGT LWWHAHSKWLRATVYGALIIYPKLGSPYPFPMPRREIPILLGEWWDRNPMDVLRQAIFTGAAPNVSDAYTINGQPGDFYRCSKQETAVFPVQSGETILLR IINAAMNQELFFSVANHKLTVVAVDAAYTKPFTTNVIMIAPGQTTNVLLTADQPPARYYMAAHAYNTANAAFDNTTTTAILEYKSAPCNAKKGKSSSSTP VFAQLPGFNDTATATAFTSSLRSPHKVHVPIT | |||
| Full Sequence |
|---|
| Protein Sequence Length: 432 Download |
| MESHNFSVNS VFSCLFFAIL FSFGSLASFA VAETHYHEFV IEERPVKRLC KTHGRITVNG 60 QFPGPTLEVR DGDSLVITAI NKAQYNVTLH WHGIRQLRNP WADGPEYITQ CPILPGRSYT 120 YRFTIINQEG TLWWHAHSKW LRATVYGALI IYPKLGSPYP FPMPRREIPI LLGEWWDRNP 180 MDVLRQAIFT GAAPNVSDAY TINGQPGDFY RCSKQETAVF PVQSGETILL RIINAAMNQE 240 LFFSVANHKL TVVAVDAAYT KPFTTNVIMI APGQTTNVLL TADQPPARYY MAAHAYNTAN 300 AAFDNTTTTA ILEYKSAPCN AKKGKSSSST PVFAQLPGFN DTATATAFTS SLRSPHKVHV 360 PITIDXXXXX XXXXXXXXXX XXXXXXXXXX XXXXXXXXXX XXXXXXXXXX XXXXXXXXXX 420 XXXXXXXXLH R* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02604 | PLN02604 | 9.0e-38 | 16 | 288 | 296 | + oxidoreductase | ||
| pfam00394 | Cu-oxidase | 5.0e-38 | 166 | 286 | 125 | + Multicopper oxidase. Many of the proteins in this family contain multiple similar copies of this plastocyanin-like domain. | ||
| TIGR03388 | ascorbase | 1.0e-46 | 50 | 361 | 337 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. | ||
| pfam07732 | Cu-oxidase_3 | 2.0e-52 | 40 | 154 | 117 | + Multicopper oxidase. This entry contains many divergent copper oxidase-like domains that are not recognised by the pfam00394 model. | ||
| TIGR03389 | laccase | 6.0e-163 | 32 | 365 | 338 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005507 | copper ion binding |
| GO:0016491 | oxidoreductase activity |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | NP_196330.3 | 0 | 16 | 365 | 7 | 355 | LAC13 (laccase 13); laccase [Arabidopsis thaliana] |
| RefSeq | XP_002277722.1 | 0 | 27 | 365 | 27 | 363 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002319955.1 | 0 | 1 | 365 | 1 | 363 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002326089.1 | 0 | 1 | 365 | 1 | 363 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002516345.1 | 0 | 1 | 365 | 1 | 364 | laccase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1asq_B | 7e-37 | 50 | 293 | 19 | 279 | X Chain X, Crystal Structure Of The Full-Length Autotransporter Esta From Pseudomonas Aeruginosa |
| PDB | 1asq_A | 7e-37 | 50 | 293 | 19 | 279 | X Chain X, Crystal Structure Of The Full-Length Autotransporter Esta From Pseudomonas Aeruginosa |
| PDB | 1asp_B | 7e-37 | 50 | 293 | 19 | 279 | X Chain X, Crystal Structure Of The Full-Length Autotransporter Esta From Pseudomonas Aeruginosa |
| PDB | 1asp_A | 7e-37 | 50 | 293 | 19 | 279 | X Chain X, Crystal Structure Of The Full-Length Autotransporter Esta From Pseudomonas Aeruginosa |
| PDB | 1aso_B | 7e-37 | 50 | 293 | 19 | 279 | X Chain X, Crystal Structure Of The Full-Length Autotransporter Esta From Pseudomonas Aeruginosa |
| Transmembrane Domains | ||||
|---|---|---|---|---|
 | ||||
| Start | End | |||
| 7 | 29 | |||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| BX839990 | 308 | 12 | 318 | 0 |
| DV141921 | 247 | 13 | 259 | 0 |
| DY812360 | 277 | 12 | 288 | 0 |
| EX658195 | 279 | 10 | 288 | 0 |
| DY812253 | 273 | 12 | 284 | 0 |
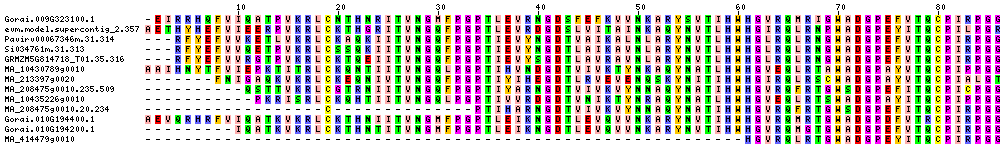
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |