You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000503_00834
You are here: Home > Sequence: MGYG000000503_00834
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
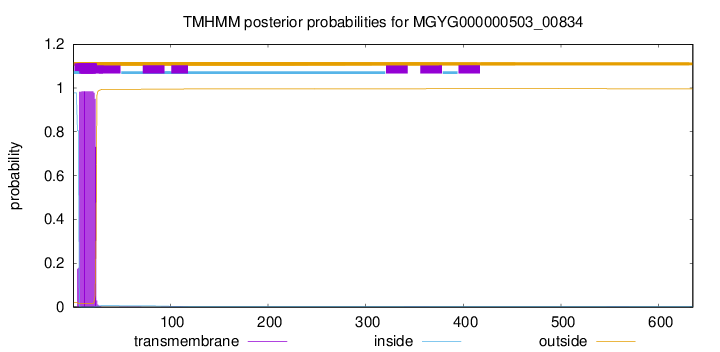
TMHMM annotations
Basic Information help
| Species | UMGS1585 sp900766355 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Cyanobacteria; Vampirovibrionia; Gastranaerophilales; Gastranaerophilaceae; UMGS1585; UMGS1585 sp900766355 | |||||||||||
| CAZyme ID | MGYG000000503_00834 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 88492; End: 90399 Strand: + | |||||||||||
Full Sequence Download help
| MKKDKIYLTI LLAIVCLISA IAVNKKLSET QDMNKYKQAI TMYQAEDYQN AYYNFSAVSE | 60 |
| FSTLKQAALF RQARCATTLG DTHTAIRNYY ILLKRFPNSP LYAISEYNLA VLLYEAQEYH | 120 |
| SAKRHFVHIY KHFKDKEVSI ASEYYLGMLE NKPELLLDYI KQSPNGRFSK SAIKTLIENN | 180 |
| TKLSNADNLV IANSYLANEN YNEALAYYQK TSLKDSWTGY AKTLYKLGKY EEAKKITIKG | 240 |
| LSIYSSFVDA NEIYDVIDCF VSLSDSKLTT IKYLLSVNPK AKGADYLMFL LAKYSPSAQT | 300 |
| SYIYEQLYTR FPDGQFSGEA LYKTFYSKIN QRKYKEAIKL GKLHLAKFGN TNSAPAVMYW | 360 |
| IAKVYEKQRM NDMARSYYKG VLSKYPDSYY AMRANAKLNK NHTMFTKNSI TNKPVVFPIH | 420 |
| NKSEADMAVK LAKLGDYDFV QELYKNDGFV QSWIEYEKGN YTNSALLARE AMDKLEVKPN | 480 |
| FKDTRWRLVY PVHYYEYIKK YSNEENPIII LSIIKEESHF NPKVTSPVGA GGLMQLMPAT | 540 |
| ANEIAKGYGI SSNLYSPSSN IHLGCLYYAK IKRTLNNKDI SAIMAYNGGW ASVMNWKEQL | 600 |
| EYVDMDDFIE KIPYPETQDY IKKVLKSYWN YCNIY | 635 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 497 | 630 | 1e-26 | 0.8740740740740741 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd13401 | Slt70-like | 1.97e-48 | 490 | 631 | 2 | 149 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| cd16896 | LT_Slt70-like | 3.09e-42 | 491 | 628 | 1 | 144 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd00254 | LT-like | 2.03e-30 | 507 | 629 | 1 | 111 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| PRK11619 | PRK11619 | 8.95e-24 | 511 | 625 | 498 | 618 | lytic murein transglycosylase; Provisional |
| pfam01464 | SLT | 7.96e-22 | 496 | 599 | 3 | 105 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOR39013.1 | 4.43e-141 | 22 | 635 | 28 | 657 |
| BAQ62516.1 | 3.63e-43 | 60 | 635 | 117 | 711 |
| ALJ69570.1 | 3.06e-39 | 149 | 635 | 181 | 702 |
| QHV01864.1 | 3.39e-39 | 149 | 635 | 197 | 718 |
| BAL33774.1 | 5.05e-39 | 149 | 635 | 278 | 799 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1SLY_A | 9.60e-20 | 462 | 631 | 414 | 600 | ComplexOf The 70-Kda Soluble Lytic Transglycosylase With Bulgecin A [Escherichia coli] |
| 1QSA_A | 1.69e-19 | 462 | 631 | 414 | 600 | CrystalStructure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.65 Angstroms Resolution [Escherichia coli],1QTE_A Crystal Structure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.90 A Resolution In Complex With A 1,6- Anhydromurotripeptide [Escherichia coli] |
| 5O1J_A | 5.91e-18 | 354 | 627 | 275 | 555 | Lytictransglycosylase in action [Neisseria meningitidis MC58] |
| 5MPQ_A | 3.16e-17 | 506 | 627 | 430 | 551 | BulgecinA: The key to a broad-spectrum inhibitor that targets lytic transglycosylases [Neisseria meningitidis] |
| 6FPN_B | 3.24e-17 | 506 | 627 | 440 | 561 | Lytictransglycosylase in action [Neisseria meningitidis MC58] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0AGC3 | 9.82e-19 | 462 | 631 | 441 | 627 | Soluble lytic murein transglycosylase OS=Escherichia coli (strain K12) OX=83333 GN=slt PE=1 SV=1 |
| P0AGC4 | 9.82e-19 | 462 | 631 | 441 | 627 | Soluble lytic murein transglycosylase OS=Escherichia coli O157:H7 OX=83334 GN=slt PE=3 SV=1 |
| P39434 | 1.52e-16 | 462 | 625 | 441 | 619 | Soluble lytic murein transglycosylase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=slt PE=3 SV=2 |
| O31976 | 7.14e-12 | 490 | 628 | 1418 | 1541 | SPbeta prophage-derived uncharacterized transglycosylase YomI OS=Bacillus subtilis (strain 168) OX=224308 GN=yomI PE=3 SV=2 |
| O64046 | 7.14e-12 | 490 | 628 | 1418 | 1541 | Probable tape measure protein OS=Bacillus phage SPbeta OX=66797 GN=yomI PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
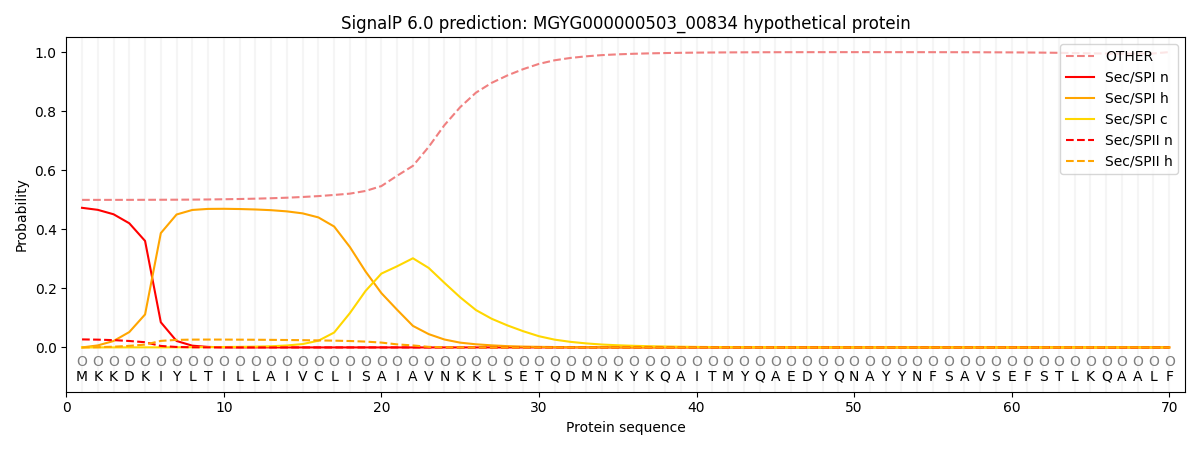
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.516395 | 0.452458 | 0.028659 | 0.000679 | 0.000481 | 0.001320 |