You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002331_00158
You are here: Home > Sequence: MGYG000002331_00158
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Vibrio parahaemolyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Vibrionaceae; Vibrio; Vibrio parahaemolyticus | |||||||||||
| CAZyme ID | MGYG000002331_00158 | |||||||||||
| CAZy Family | GH63 | |||||||||||
| CAZyme Description | Glucosidase YgjK | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 152050; End: 154740 Strand: + | |||||||||||
Full Sequence Download help
| MFKKSTLAIL IGATISLVGC GSNDKTEIRL PQAPTSEFTN VIDRTGNPTY LRDYDSYSNL | 60 |
| KYNALTDNGA WHGHLLPEDA AGYGAFGGIM QVTQEYAHFM SGQTFDKLTL TDKVSGQTYD | 120 |
| LSNAEAKVYS IPGALVQVLD LDDIHVQMVL RFVTDRSSLV ETLITNKTDR DMNLALEWDG | 180 |
| ELVKYALSTR KDQTVAQYYP DYKRQISTIE NGIKINFSEM KDNWAIRNDR DAEFRITRSL | 240 |
| PTKTFTNETS FSSTAEMSLP AEQTSAVYTA YSNLHNAKEV QSESLKLQDV LANPTQYMEA | 300 |
| SKARWDGYLA NGLINKEATA DQARVAVKAM ETLNGNWRSA AGDIEQATVT PSVTARWFSG | 360 |
| NLTWPWDSWK QAYAMAHFNP DVAMDNIRTV FQEQVQADDK IRPQDKGYLL DVLTYTKPVS | 420 |
| RGGAGSENWN ERNTKPSLAA WSVMEVYHAL VNQFDRPEDA QKFIDEMYPK LVAYHDWWLS | 480 |
| NRDHNQNGVP EYGAAVDPAH NTEEGVMYVW VETRDPNFTT IIPAEDIIEQ NGNSYKVKGM | 540 |
| ASYNKILDEV DYMTIHVGAQ EAAGWESGMD NAARFGFIYN IHNMNSQAED ISNPDRNVDQ | 600 |
| LGRYAASAYG FDNSIKLEGE EWVYSNTSAE NLAKLDLAKK DWEVRFAENR TNNNAADGEL | 660 |
| LGFSMLQESV DQASYMYSDN KYLAEMAKLV SDGVEGKDRA QEFLNGAEKI KTYINQCMFD | 720 |
| ESTGFFYDIH LNVAEDGTTP APLANGCAGL PIVERGRGPE GWSPLFNGAA TQEHADKVIA | 780 |
| VMRDQDEFNT PDKFQGTGVP LPTASQTNPA YGKDVYWRGR VWLDQVYFGF RAMENYGYKE | 840 |
| EAIAMANELF NNAEGMTGDM PIRENYNPET GAVQGASNFS WSAAHLYMMY NGFFKN | 896 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH63 | 2 | 895 | 2e-262 | 0.9982456140350877 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10137 | PRK10137 | 0.0 | 2 | 895 | 1 | 785 | alpha-glucosidase; Provisional |
| pfam03200 | Glyco_hydro_63 | 1.53e-07 | 670 | 889 | 252 | 492 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| pfam01204 | Trehalase | 5.16e-04 | 670 | 879 | 305 | 486 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNE55225.1 | 0.0 | 1 | 896 | 1 | 896 |
| QQD00876.1 | 0.0 | 1 | 896 | 1 | 896 |
| QWL01953.1 | 0.0 | 1 | 896 | 1 | 896 |
| AGB11467.1 | 0.0 | 1 | 896 | 1 | 896 |
| BCB44003.1 | 0.0 | 1 | 896 | 1 | 896 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3W7S_A | 1.36e-273 | 38 | 895 | 5 | 758 | Escherichiacoli K12 YgjK complexed with glucose [Escherichia coli K-12],3W7S_B Escherichia coli K12 YgjK complexed with glucose [Escherichia coli K-12],3W7T_A Escherichia coli K12 YgjK complexed with mannose [Escherichia coli K-12],3W7T_B Escherichia coli K12 YgjK complexed with mannose [Escherichia coli K-12],3W7U_A Escherichia coli K12 YgjK complexed with galactose [Escherichia coli K-12],3W7U_B Escherichia coli K12 YgjK complexed with galactose [Escherichia coli K-12] |
| 3W7X_A | 7.77e-273 | 38 | 895 | 5 | 758 | Crystalstructure of E. coli YgjK D324N complexed with melibiose [Escherichia coli K-12],3W7X_B Crystal structure of E. coli YgjK D324N complexed with melibiose [Escherichia coli K-12],5CA3_A Crystal structure of the glycosynthase mutant D324N of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12],5CA3_B Crystal structure of the glycosynthase mutant D324N of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12] |
| 3W7W_A | 1.10e-272 | 38 | 895 | 5 | 758 | Crystalstructure of E. coli YgjK E727A complexed with 2-O-alpha-D-glucopyranosyl-alpha-D-galactopyranose [Escherichia coli K-12],3W7W_B Crystal structure of E. coli YgjK E727A complexed with 2-O-alpha-D-glucopyranosyl-alpha-D-galactopyranose [Escherichia coli K-12],5GW7_A Crystal structure of the glycosynthase mutant E727A of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12],5GW7_B Crystal structure of the glycosynthase mutant E727A of Escherichia coli GH63 glycosidase in complex with glucose and lactose [Escherichia coli K-12] |
| 3D3I_A | 4.29e-262 | 33 | 895 | 1 | 759 | Crystalstructural of Escherichia coli K12 YgjK, a glucosidase belonging to glycoside hydrolase family 63 [Escherichia coli K-12],3D3I_B Crystal structural of Escherichia coli K12 YgjK, a glucosidase belonging to glycoside hydrolase family 63 [Escherichia coli K-12] |
| 7PQQ_B | 7.35e-151 | 19 | 512 | 312 | 786 | ChainB, Anti-RON nanobody,Megabody 91,Glucosidase YgjK [Lama glama] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P42592 | 8.27e-273 | 33 | 895 | 23 | 781 | Glucosidase YgjK OS=Escherichia coli (strain K12) OX=83333 GN=ygjK PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
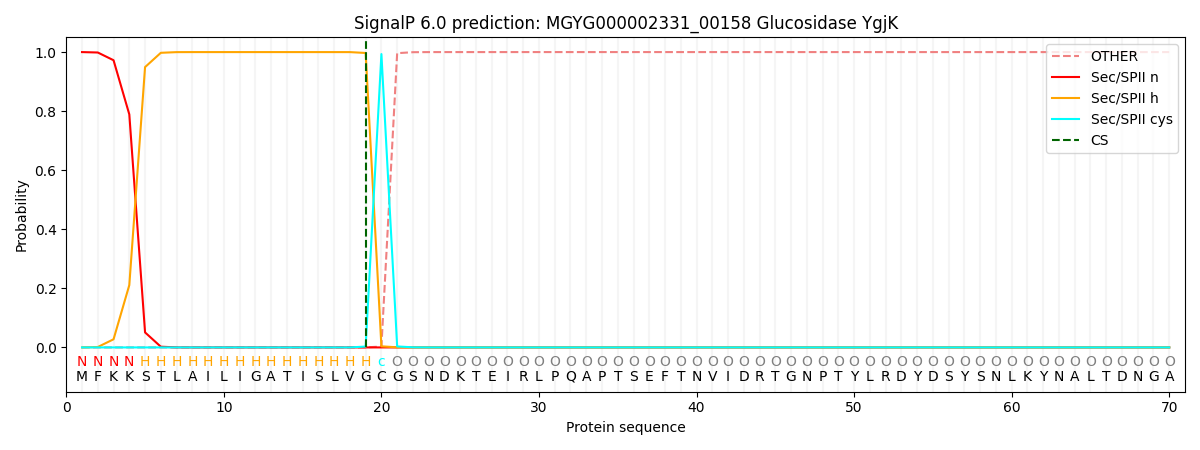
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000050 | 0.000000 | 0.000000 | 0.000000 |
