You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004865_00263
You are here: Home > Sequence: MGYG000004865_00263
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Spirochaetota; Spirochaetia; Sphaerochaetales; Sphaerochaetaceae; RUG023; | |||||||||||
| CAZyme ID | MGYG000004865_00263 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4427; End: 5752 Strand: - | |||||||||||
Full Sequence Download help
| MRKALAILVL CAVGAALHGQ PTQEVRLDEK HPSLELVGYG SDTKGGMGGT IIKVTNLHAD | 60 |
| GEGSFKAALE AEGPRIIVFE VGGIIDLEKQ QIKVKNPYVT IAGQTAPSPG ITCIRGGLTL | 120 |
| DTHDIVMQHI RFRMGDAGAE PGSGYEPEIS TSAHATAYNI VVDHCSVSWG IDENLSVSGT | 180 |
| RGGDGAKSPH DVTLSYNFIS ECLRYSVHSK TKSKHMDHSM GTLVMDDVKR VAIIGNYYAH | 240 |
| NSERNPWFKA NSSGVVVNNL IYDPGTWAIR LSWIPKEWKN VKDGKPGTPV VSIVGNYEKE | 300 |
| GPSTKVVAMV GSNYPENHES GYLEDNICVK ADGSQGKVTD PSIPIGQLDQ RPIWIDGLKV | 360 |
| LPAKEVPDYI LKWAGARPAD RDQTDLRLID EFTKGTGKII DSQDEVGGYP TATPTYRKLD | 420 |
| VPDEHVEEWL EGFTKEVMGL D | 441 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 82 | 269 | 2.3e-49 | 0.907103825136612 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3866 | PelB | 1.17e-06 | 43 | 246 | 48 | 234 | Pectate lyase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QYK69250.1 | 6.84e-149 | 37 | 437 | 43 | 433 |
| ALA44843.1 | 1.26e-148 | 37 | 437 | 20 | 410 |
| APB74413.1 | 2.76e-148 | 37 | 437 | 43 | 433 |
| AIW40090.1 | 2.76e-148 | 37 | 437 | 43 | 433 |
| ALP35378.1 | 7.85e-148 | 7 | 437 | 19 | 433 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5B297 | 1.16e-32 | 5 | 424 | 8 | 377 | Probable pectate lyase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyC PE=3 SV=1 |
| B8NQQ7 | 2.93e-32 | 38 | 441 | 28 | 399 | Probable pectate lyase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyC PE=3 SV=1 |
| Q2UB83 | 5.14e-31 | 38 | 441 | 28 | 399 | Probable pectate lyase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=plyC PE=3 SV=1 |
| A1DPF0 | 1.86e-30 | 38 | 408 | 29 | 361 | Probable pectate lyase C OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyC PE=3 SV=1 |
| Q0CLG7 | 3.14e-29 | 38 | 408 | 28 | 360 | Probable pectate lyase C OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
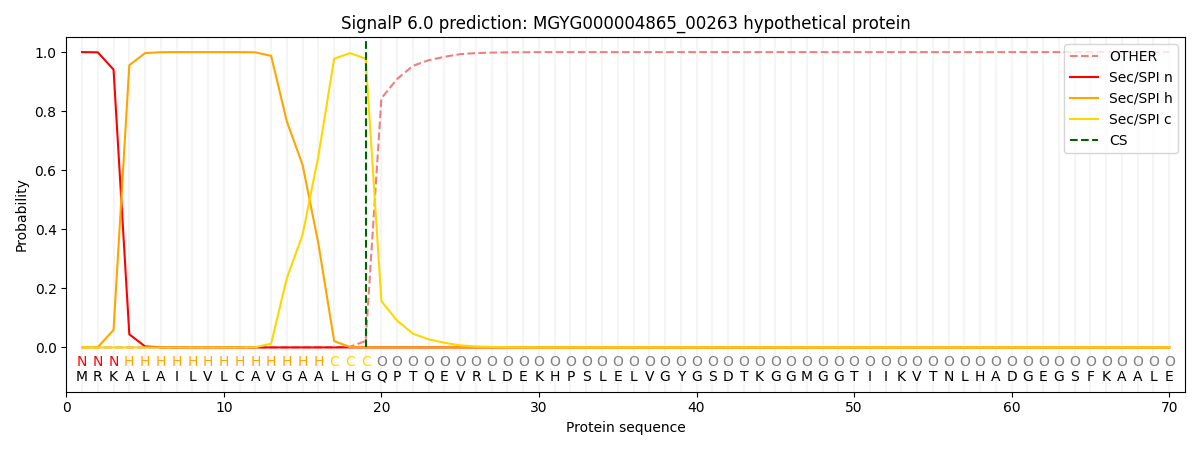
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000272 | 0.999036 | 0.000200 | 0.000166 | 0.000172 | 0.000148 |
