You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000012_03421
You are here: Home > Sequence: MGYG000000012_03421
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacillus subtilis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales; Bacillaceae; Bacillus; Bacillus subtilis | |||||||||||
| CAZyme ID | MGYG000000012_03421 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | Pectate lyase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 215537; End: 216799 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 134 | 345 | 4.6e-101 | 0.9906103286384976 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3866 | PelB | 2.20e-120 | 1 | 420 | 1 | 345 | Pectate lyase [Carbohydrate transport and metabolism]. |
| smart00656 | Amb_all | 2.98e-63 | 144 | 350 | 12 | 190 | Amb_all domain. |
| pfam00544 | Pec_lyase_C | 1.76e-40 | 130 | 345 | 16 | 210 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRZ93715.1 | 6.02e-308 | 1 | 420 | 1 | 420 |
| AFP43212.1 | 6.02e-308 | 1 | 420 | 1 | 420 |
| ASB60052.1 | 7.03e-307 | 1 | 420 | 1 | 420 |
| QGI50986.1 | 9.98e-307 | 1 | 420 | 1 | 420 |
| AFQ56675.1 | 9.98e-307 | 1 | 420 | 1 | 420 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1BN8_A | 3.65e-308 | 1 | 420 | 1 | 420 | BacillusSubtilis Pectate Lyase [Bacillus subtilis] |
| 2BSP_A | 1.05e-307 | 1 | 420 | 1 | 420 | ChainA, PROTEIN (PECTATE LYASE) [Bacillus subtilis] |
| 5AMV_A | 2.87e-293 | 22 | 420 | 1 | 399 | Structuralinsights into the loss of catalytic competence in pectate lyase at low pH [Bacillus subtilis],5X2I_A Polygalacturonate Lyase by Fusing with a Self-assembling Amphipathic Peptide [Bacillus subtilis subsp. subtilis str. 168] |
| 2NZM_A | 2.36e-292 | 22 | 420 | 1 | 399 | ChainA, Pectate lyase [Bacillus subtilis],2O04_A Chain A, Pectate lyase [Bacillus subtilis],2O0V_A Chain A, Pectate lyase [Bacillus subtilis],2O0W_A Chain A, Pectate lyase [Bacillus subtilis],2O17_A Chain A, Pectate lyase [Bacillus subtilis],2O1D_A Chain A, Pectate lyase [Bacillus subtilis] |
| 3KRG_A | 6.47e-290 | 22 | 420 | 1 | 399 | ChainA, Pectate lyase [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39116 | 2.00e-307 | 1 | 420 | 1 | 420 | Pectate lyase OS=Bacillus subtilis (strain 168) OX=224308 GN=pel PE=1 SV=1 |
| Q60140 | 5.04e-99 | 6 | 419 | 12 | 379 | Pectate lyase OS=Pseudomonas viridiflava OX=33069 GN=pel PE=3 SV=1 |
| Q59671 | 7.27e-96 | 9 | 419 | 19 | 380 | Pectate lyase OS=Pseudomonas fluorescens OX=294 GN=pel PE=3 SV=1 |
| Q56806 | 9.36e-96 | 7 | 419 | 10 | 377 | Pectate lyase OS=Xanthomonas campestris pv. malvacearum OX=86040 PE=3 SV=1 |
| P72242 | 7.95e-95 | 34 | 419 | 39 | 379 | Pectate lyase OS=Pseudomonas amygdali pv. lachrymans OX=53707 GN=pelP PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
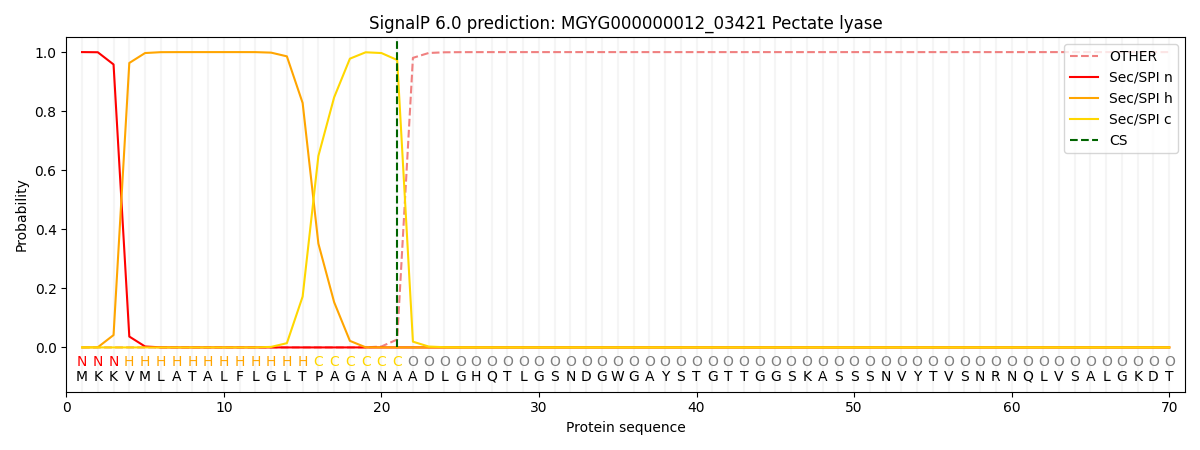
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000212 | 0.999141 | 0.000161 | 0.000164 | 0.000155 | 0.000138 |
