You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000012_03536
You are here: Home > Sequence: MGYG000000012_03536
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacillus subtilis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales; Bacillaceae; Bacillus; Bacillus subtilis | |||||||||||
| CAZyme ID | MGYG000000012_03536 | |||||||||||
| CAZy Family | GH26 | |||||||||||
| CAZyme Description | Mannan endo-1,4-beta-mannosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1020; End: 2108 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH26 | 32 | 350 | 3.1e-93 | 0.9966996699669967 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02156 | Glyco_hydro_26 | 3.47e-165 | 32 | 350 | 1 | 311 | Glycosyl hydrolase family 26. |
| COG4124 | ManB2 | 5.69e-121 | 1 | 362 | 1 | 355 | Beta-mannanase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRZ93549.1 | 3.91e-270 | 1 | 362 | 1 | 362 |
| AFI27190.1 | 1.12e-269 | 1 | 362 | 1 | 362 |
| BBA72035.1 | 1.12e-269 | 1 | 362 | 1 | 362 |
| ASB59884.1 | 9.22e-269 | 1 | 362 | 1 | 362 |
| AUZ37462.1 | 1.31e-268 | 1 | 362 | 1 | 362 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3CBW_A | 6.24e-257 | 20 | 362 | 3 | 345 | ChainA, YdhT protein [Bacillus subtilis],3CBW_B Chain B, YdhT protein [Bacillus subtilis] |
| 2QHA_A | 1.32e-253 | 27 | 362 | 1 | 336 | FromStructure to Function: Insights into the Catalytic Substrate Specificity and Thermostability Displayed by Bacillus subtilis mannanase BCman [Bacillus subtilis],2QHA_B From Structure to Function: Insights into the Catalytic Substrate Specificity and Thermostability Displayed by Bacillus subtilis mannanase BCman [Bacillus subtilis] |
| 2WHK_A | 4.12e-251 | 27 | 362 | 1 | 336 | Structureof Bacillus subtilis mannanase man26 [Bacillus subtilis] |
| 7EET_A | 1.71e-119 | 26 | 351 | 7 | 345 | ChainA, Mannanase KMAN [Klebsiella oxytoca] |
| 6Q75_A | 3.55e-29 | 31 | 351 | 23 | 327 | Thestructure of GH26A from Muricauda sp. MAR_2010_75 [Muricauda sp. MAR_2010_75],6Q75_B The structure of GH26A from Muricauda sp. MAR_2010_75 [Muricauda sp. MAR_2010_75] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O05512 | 1.19e-265 | 1 | 362 | 1 | 362 | Mannan endo-1,4-beta-mannosidase OS=Bacillus subtilis (strain 168) OX=224308 GN=gmuG PE=1 SV=2 |
| P55278 | 1.69e-205 | 1 | 362 | 1 | 360 | Mannan endo-1,4-beta-mannosidase OS=Bacillus subtilis OX=1423 PE=3 SV=1 |
| P16699 | 3.07e-117 | 33 | 361 | 36 | 365 | Mannan endo-1,4-beta-mannosidase A and B OS=Caldalkalibacillus mannanilyticus (strain DSM 16130 / CIP 109019 / JCM 10596 / AM-001) OX=1236954 PE=1 SV=1 |
| Q5AWB7 | 1.19e-18 | 9 | 294 | 6 | 298 | Probable mannan endo-1,4-beta-mannosidase E OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=manE PE=3 SV=1 |
| A1A278 | 1.91e-18 | 35 | 324 | 46 | 358 | Mannan endo-1,4-beta-mannosidase OS=Bifidobacterium adolescentis (strain ATCC 15703 / DSM 20083 / NCTC 11814 / E194a) OX=367928 GN=BAD_1030 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
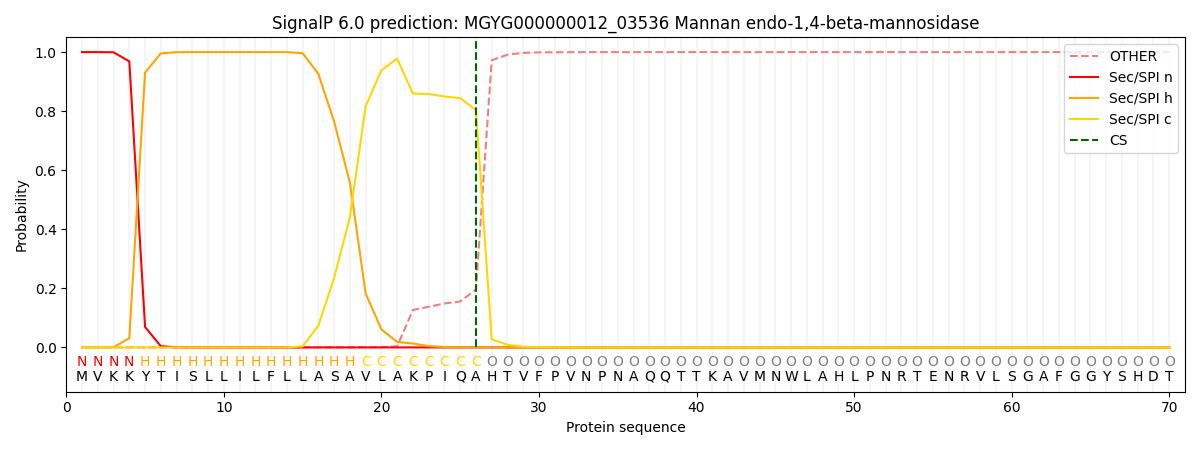
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000206 | 0.999235 | 0.000145 | 0.000141 | 0.000127 | 0.000122 |