You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000013_02451
You are here: Home > Sequence: MGYG000000013_02451
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp902362375 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp902362375 | |||||||||||
| CAZyme ID | MGYG000000013_02451 | |||||||||||
| CAZy Family | CBM85 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 478479; End: 480260 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 247 | 533 | 8.8e-64 | 0.9504950495049505 |
| CBM85 | 51 | 180 | 3.3e-26 | 0.9772727272727273 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 9.30e-43 | 287 | 534 | 1 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 1.48e-42 | 251 | 516 | 13 | 283 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 4.64e-36 | 246 | 525 | 33 | 318 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| pfam13620 | CarboxypepD_reg | 0.005 | 222 | 257 | 2 | 37 | Carboxypeptidase regulatory-like domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AEE54613.1 | 3.56e-166 | 2 | 413 | 8 | 420 |
| CCW34445.1 | 4.95e-127 | 25 | 583 | 32 | 578 |
| QYM77625.1 | 6.40e-125 | 9 | 577 | 7 | 583 |
| AHF89713.1 | 2.64e-124 | 27 | 575 | 64 | 616 |
| QDU72038.1 | 6.30e-123 | 86 | 591 | 104 | 603 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7D88_A | 1.68e-53 | 200 | 583 | 38 | 412 | ChainA, Beta-xylanase [Bacillus sp. (in: Bacteria)] |
| 7D89_A | 8.59e-52 | 200 | 583 | 38 | 412 | ChainA, Beta-xylanase [Bacillus sp. (in: Bacteria)] |
| 6FHE_A | 4.69e-27 | 248 | 522 | 21 | 320 | Highlyactive enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
| 3NIY_A | 7.55e-26 | 265 | 541 | 45 | 339 | Crystalstructure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NIY_B Crystal structure of native xylanase 10B from Thermotoga petrophila RKU-1 [Thermotoga petrophila RKU-1],3NJ3_A Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1],3NJ3_B Crystal structure of xylanase 10B from Thermotoga petrophila RKU-1 in complex with xylobiose [Thermotoga petrophila RKU-1] |
| 1VBR_A | 2.77e-25 | 265 | 541 | 29 | 323 | Crystalstructure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBR_B Crystal structure of complex xylanase 10B from Thermotoga maritima with xylobiose [Thermotoga maritima],1VBU_A Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima],1VBU_B Crystal structure of native xylanase 10B from Thermotoga maritima [Thermotoga maritima] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A3DH97 | 1.60e-51 | 206 | 573 | 380 | 736 | Anti-sigma-I factor RsgI6 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=rsgI6 PE=1 SV=1 |
| A0A1P8AWH8 | 2.36e-41 | 213 | 583 | 567 | 930 | Endo-1,4-beta-xylanase 1 OS=Arabidopsis thaliana OX=3702 GN=XYN1 PE=1 SV=1 |
| O80596 | 4.28e-37 | 218 | 592 | 694 | 1060 | Endo-1,4-beta-xylanase 2 OS=Arabidopsis thaliana OX=3702 GN=XYN2 PE=3 SV=1 |
| F4JG10 | 9.59e-33 | 93 | 591 | 280 | 745 | Endo-1,4-beta-xylanase 3 OS=Arabidopsis thaliana OX=3702 GN=XYN3 PE=2 SV=1 |
| Q84WT5 | 1.38e-30 | 206 | 571 | 173 | 540 | Endo-1,4-beta-xylanase 5-like OS=Arabidopsis thaliana OX=3702 GN=At4g33820 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
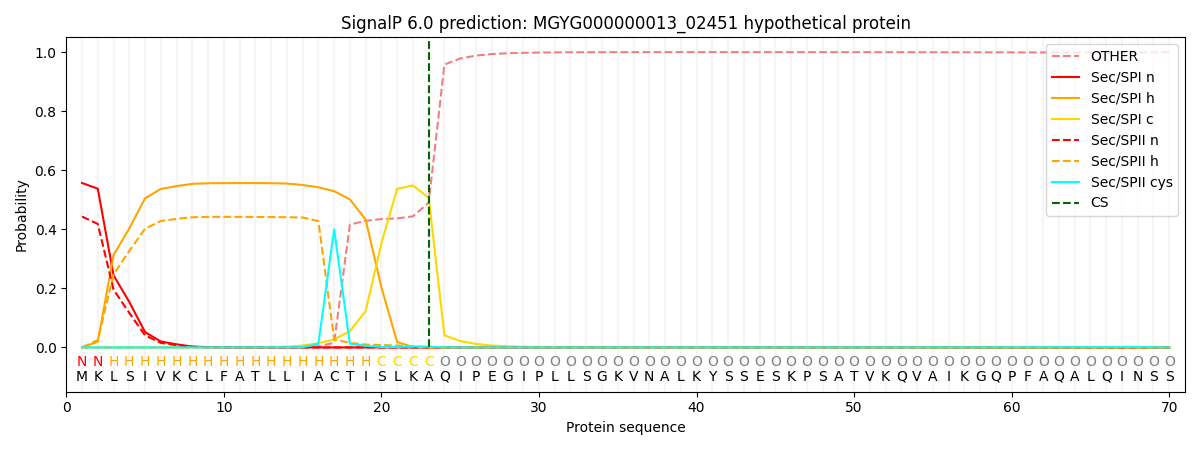
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000488 | 0.547337 | 0.451520 | 0.000214 | 0.000224 | 0.000196 |
