You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000013_02943
You are here: Home > Sequence: MGYG000000013_02943
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp902362375 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp902362375 | |||||||||||
| CAZyme ID | MGYG000000013_02943 | |||||||||||
| CAZy Family | CE8 | |||||||||||
| CAZyme Description | Pectinesterase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 72515; End: 73486 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE8 | 33 | 314 | 1.8e-107 | 0.9895833333333334 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01095 | Pectinesterase | 1.76e-83 | 34 | 311 | 3 | 293 | Pectinesterase. |
| PLN02773 | PLN02773 | 2.57e-83 | 27 | 307 | 1 | 290 | pectinesterase |
| PLN02432 | PLN02432 | 3.15e-74 | 33 | 306 | 13 | 283 | putative pectinesterase |
| PLN02708 | PLN02708 | 4.51e-70 | 35 | 321 | 245 | 550 | Probable pectinesterase/pectinesterase inhibitor |
| PLN02682 | PLN02682 | 1.59e-69 | 32 | 303 | 70 | 351 | pectinesterase family protein |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNL41079.1 | 2.77e-223 | 1 | 323 | 1 | 323 |
| QUR41788.1 | 2.54e-219 | 1 | 323 | 1 | 323 |
| ALJ48951.1 | 5.13e-219 | 1 | 323 | 1 | 323 |
| QGT73499.1 | 5.13e-219 | 1 | 323 | 1 | 323 |
| QRQ55762.1 | 5.13e-219 | 1 | 323 | 1 | 323 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1GQ8_A | 1.06e-42 | 33 | 309 | 9 | 298 | Pectinmethylesterase from Carrot [Daucus carota] |
| 1XG2_A | 2.25e-40 | 34 | 319 | 6 | 303 | ChainA, Pectinesterase 1 [Solanum lycopersicum] |
| 5C1E_A | 4.43e-29 | 33 | 292 | 11 | 271 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Penultimately Deglycosylated Form (N-acetylglucosamine Stub at Asn84) [Aspergillus niger ATCC 1015] |
| 5C1C_A | 2.30e-28 | 33 | 292 | 11 | 271 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Deglycosylated Form [Aspergillus niger ATCC 1015] |
| 2NTB_A | 8.15e-24 | 43 | 321 | 18 | 339 | Crystalstructure of pectin methylesterase in complex with hexasaccharide V [Dickeya dadantii 3937],2NTB_B Crystal structure of pectin methylesterase in complex with hexasaccharide V [Dickeya dadantii 3937],2NTP_A Crystal structure of pectin methylesterase in complex with hexasaccharide VI [Dickeya dadantii 3937],2NTP_B Crystal structure of pectin methylesterase in complex with hexasaccharide VI [Dickeya dadantii 3937],2NTQ_A Crystal structure of pectin methylesterase in complex with hexasaccharide VII [Dickeya dadantii 3937],2NTQ_B Crystal structure of pectin methylesterase in complex with hexasaccharide VII [Dickeya dadantii 3937] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9LVQ0 | 1.06e-50 | 35 | 303 | 9 | 286 | Pectinesterase 31 OS=Arabidopsis thaliana OX=3702 GN=PME31 PE=1 SV=1 |
| P41510 | 1.21e-50 | 30 | 319 | 270 | 572 | Probable pectinesterase/pectinesterase inhibitor OS=Brassica napus OX=3708 GN=BP19 PE=2 SV=1 |
| Q42608 | 1.97e-48 | 30 | 319 | 257 | 559 | Pectinesterase/pectinesterase inhibitor (Fragment) OS=Brassica campestris OX=3711 PE=2 SV=1 |
| Q8GX86 | 2.55e-48 | 30 | 323 | 253 | 557 | Probable pectinesterase/pectinesterase inhibitor 21 OS=Arabidopsis thaliana OX=3702 GN=PME21 PE=2 SV=2 |
| O49298 | 2.94e-48 | 30 | 319 | 248 | 542 | Probable pectinesterase/pectinesterase inhibitor 6 OS=Arabidopsis thaliana OX=3702 GN=PME6 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
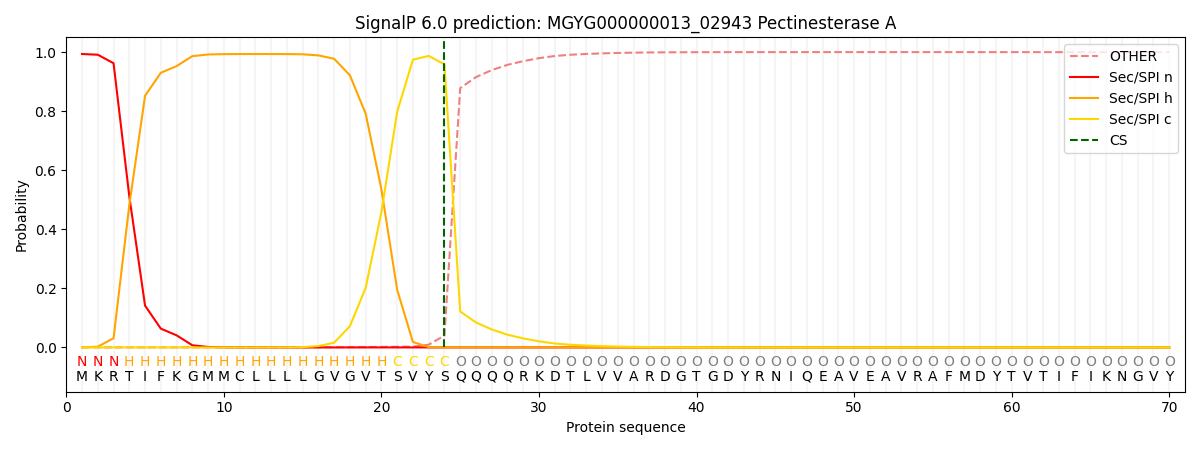
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001138 | 0.990395 | 0.007640 | 0.000284 | 0.000264 | 0.000255 |
