You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000013_03391
You are here: Home > Sequence: MGYG000000013_03391
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
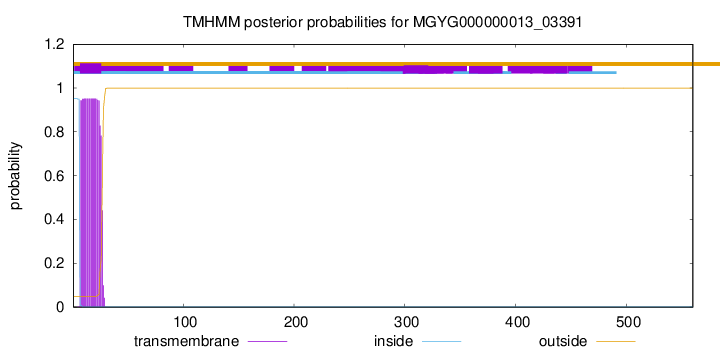
TMHMM annotations
Basic Information help
| Species | Bacteroides sp902362375 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp902362375 | |||||||||||
| CAZyme ID | MGYG000000013_03391 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 200037; End: 201719 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 134 | 324 | 1.6e-65 | 0.994535519125683 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam18884 | TSP3_bac | 2.95e-04 | 492 | 513 | 1 | 22 | Bacterial TSP3 repeat. This entry contains a novel bacterial thrombospondin type 3 repeat which differs from the typical consensus by containing a glutamate in place of one of the calcium binding aspartate residues. |
| cd16230 | EFh_CREC_RCN3 | 6.14e-04 | 465 | 532 | 115 | 183 | EF-hand, calcium binding motif, found in reticulocalbin-3 (RCN-3). RCN-3, also termed EF-hand calcium-binding protein RLP49, is a putative six EF-hand Ca2+-binding protein that contains five RXXR (X is any amino acid) motifs and a C-terminal ER retrieval signal His-Asp-Glu-Leu (HDEL) tetrapeptide. The RXXR motif represents the target sequence of subtilisin-like proprotein convertases (SPCs). RCN-3 is specifically bound to the paired basic amino-acid-cleaving enzyme-4 (PACE4) precursor protein and plays an important role in the biosynthesis of PACE4. |
| smart00656 | Amb_all | 0.003 | 149 | 250 | 17 | 111 | Amb_all domain. |
| COG3866 | PelB | 0.006 | 81 | 270 | 27 | 209 | Pectate lyase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNL37697.1 | 0.0 | 1 | 560 | 1 | 560 |
| QUR43057.1 | 0.0 | 1 | 560 | 1 | 560 |
| CBK65670.1 | 0.0 | 1 | 560 | 1 | 560 |
| QRN01889.1 | 0.0 | 1 | 560 | 1 | 560 |
| QUT29157.1 | 0.0 | 1 | 560 | 1 | 560 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B8NQQ7 | 1.47e-46 | 80 | 511 | 18 | 391 | Probable pectate lyase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyC PE=3 SV=1 |
| Q2UB83 | 1.01e-44 | 80 | 511 | 18 | 391 | Probable pectate lyase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=plyC PE=3 SV=1 |
| Q0CLG7 | 9.82e-44 | 80 | 511 | 18 | 391 | Probable pectate lyase C OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyC PE=3 SV=1 |
| Q4WL88 | 1.91e-43 | 84 | 511 | 23 | 392 | Probable pectate lyase C OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyC PE=3 SV=1 |
| A1DPF0 | 2.65e-43 | 84 | 511 | 23 | 392 | Probable pectate lyase C OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000398 | 0.997343 | 0.001691 | 0.000198 | 0.000173 | 0.000171 |