You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000018_00025
You are here: Home > Sequence: MGYG000000018_00025
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
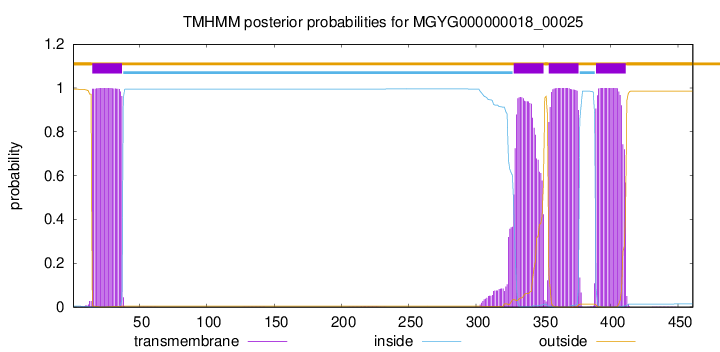
TMHMM annotations
Basic Information help
| Species | Coprococcus eutactus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Coprococcus; Coprococcus eutactus | |||||||||||
| CAZyme ID | MGYG000000018_00025 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 27335; End: 28720 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 58 | 231 | 3.1e-21 | 0.9588235294117647 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| TIGR03111 | glyc2_xrt_Gpos1 | 0.0 | 12 | 445 | 7 | 439 | putative glycosyltransferase, exosortase G-associated. Members of this protein family are probable glycosyltransferases of family 2, whose genes are near those for the exosortase homolog XrtG (TIGR03110), which is restricted to Gram-positive bacteria. Other genes in the conserved gene neighborhood include a 6-pyruvoyl tetrahydropterin synthase homolog (TIGR03112) and an uncharacterized intergral membrane protein (TIGR03766). |
| COG1215 | BcsA | 8.07e-60 | 11 | 425 | 8 | 424 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| cd06423 | CESA_like | 7.10e-36 | 58 | 246 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| PRK11204 | PRK11204 | 2.67e-29 | 1 | 384 | 3 | 367 | N-glycosyltransferase; Provisional |
| cd00761 | Glyco_tranf_GTA_type | 1.67e-23 | 59 | 210 | 2 | 153 | Glycosyltransferase family A (GT-A) includes diverse families of glycosyl transferases with a common GT-A type structural fold. Glycosyltransferases (GTs) are enzymes that synthesize oligosaccharides, polysaccharides, and glycoconjugates by transferring the sugar moiety from an activated nucleotide-sugar donor to an acceptor molecule, which may be a growing oligosaccharide, a lipid, or a protein. Based on the stereochemistry of the donor and acceptor molecules, GTs are classified as either retaining or inverting enzymes. To date, all GT structures adopt one of two possible folds, termed GT-A fold and GT-B fold. This hierarchy includes diverse families of glycosyl transferases with a common GT-A type structural fold, which has two tightly associated beta/alpha/beta domains that tend to form a continuous central sheet of at least eight beta-strands. The majority of the proteins in this superfamily are Glycosyltransferase family 2 (GT-2) proteins. But it also includes families GT-43, GT-6, GT-8, GT13 and GT-7; which are evolutionarily related to GT-2 and share structure similarities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK84035.1 | 1.24e-278 | 6 | 440 | 2 | 435 |
| QNU66959.1 | 1.35e-183 | 10 | 453 | 9 | 450 |
| AEI32443.1 | 1.61e-182 | 11 | 450 | 11 | 449 |
| AAK80527.1 | 1.61e-182 | 11 | 450 | 11 | 449 |
| ADZ21626.1 | 1.61e-182 | 11 | 450 | 11 | 449 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6YV7_B | 7.66e-13 | 52 | 268 | 40 | 248 | MannosyltransferasePcManGT from Pyrobaculum calidifontis [Pyrobaculum calidifontis JCM 11548],6YV8_B Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP and Mn2+ [Pyrobaculum calidifontis JCM 11548],6YV9_A Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP-Man and Mn2+ [Pyrobaculum calidifontis JCM 11548] |
| 6YV7_A | 7.71e-13 | 52 | 268 | 41 | 249 | MannosyltransferasePcManGT from Pyrobaculum calidifontis [Pyrobaculum calidifontis JCM 11548],6YV8_A Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP and Mn2+ [Pyrobaculum calidifontis JCM 11548],6YV9_B Mannosyltransferase PcManGT from Pyrobaculum calidifontis in complex with GDP-Man and Mn2+ [Pyrobaculum calidifontis JCM 11548] |
| 6P61_A | 4.34e-11 | 53 | 187 | 12 | 139 | Structureof a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_B Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_C Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197],6P61_D Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197) [Leptospira borgpetersenii serovar Hardjo-bovis str. JB197] |
| 5HEA_A | 2.40e-08 | 56 | 157 | 7 | 105 | CgTstructure in hexamer [Streptococcus parasanguinis FW213],5HEA_B CgT structure in hexamer [Streptococcus parasanguinis FW213],5HEA_C CgT structure in hexamer [Streptococcus parasanguinis FW213],5HEC_A CgT structure in dimer [Streptococcus parasanguinis FW213],5HEC_B CgT structure in dimer [Streptococcus parasanguinis FW213] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8XAR5 | 3.62e-24 | 53 | 301 | 74 | 307 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Escherichia coli O157:H7 OX=83334 GN=pgaC PE=3 SV=1 |
| P75905 | 3.62e-24 | 53 | 301 | 74 | 307 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Escherichia coli (strain K12) OX=83333 GN=pgaC PE=1 SV=1 |
| Q8GLC5 | 2.56e-19 | 56 | 295 | 49 | 275 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus epidermidis OX=1282 GN=icaA PE=3 SV=1 |
| Q5HCN1 | 3.45e-19 | 56 | 339 | 49 | 312 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain COL) OX=93062 GN=icaA PE=3 SV=1 |
| Q6GDD8 | 3.45e-19 | 56 | 339 | 49 | 312 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=icaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000034 | 0.000015 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |