You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000019_00553
You are here: Home > Sequence: MGYG000000019_00553
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Enterobacter cloacae_O | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Enterobacter; Enterobacter cloacae_O | |||||||||||
| CAZyme ID | MGYG000000019_00553 | |||||||||||
| CAZy Family | CBM5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 179382; End: 182105 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd04843 | Peptidases_S8_11 | 9.28e-65 | 192 | 451 | 11 | 277 | Peptidase S8 family domain, uncharacterized subfamily 11. This family is a member of the Peptidases S8 or Subtilases serine endo- and exo-peptidase clan. They have an Asp/His/Ser catalytic triad similar to that found in trypsin-like proteases, but do not share their three-dimensional structure and are not homologous to trypsin. The stability of subtilases may be enhanced by calcium, some members have been shown to bind up to 4 ions via binding sites with different affinity. Some members of this clan contain disulfide bonds. These enzymes can be intra- and extracellular, some function at extreme temperatures and pH values. |
| cd07498 | Peptidases_S8_15 | 9.46e-12 | 212 | 449 | 20 | 242 | Peptidase S8 family domain, uncharacterized subfamily 15. This family is a member of the Peptidases S8 or Subtilases serine endo- and exo-peptidase clan. They have an Asp/His/Ser catalytic triad similar to that found in trypsin-like proteases, but do not share their three-dimensional structure and are not homologous to trypsin. The stability of subtilases may be enhanced by calcium, some members have been shown to bind up to 4 ions via binding sites with different affinity. Some members of this clan contain disulfide bonds. These enzymes can be intra- and extracellular, some function at extreme temperatures and pH values. |
| cd04077 | Peptidases_S8_PCSK9_ProteinaseK_like | 2.11e-11 | 309 | 452 | 133 | 255 | Peptidase S8 family domain in ProteinaseK-like proteins. The peptidase S8 or Subtilase clan of proteases have a Asp/His/Ser catalytic triad that is not homologous to trypsin. This CD contains several members of this clan including: PCSK9 (Proprotein convertase subtilisin/kexin type 9), Proteinase_K, Proteinase_T, and other subtilisin-like serine proteases. PCSK9 posttranslationally regulates hepatic low-density lipoprotein receptors (LDLRs) by binding to LDLRs on the cell surface, leading to their degradation. The binding site of PCSK9 has been localized to the epidermal growth factor-like repeat A (EGF-A) domain of the LDLR. Characterized Proteinases K are secreted endopeptidases with a high degree of sequence conservation. Proteinases K are not substrate-specific and function in a wide variety of species in different pathways. It can hydrolyze keratin and other proteins with subtilisin-like specificity. The number of calcium-binding motifs found in these differ. Proteinase T is a novel proteinase from the fungus Tritirachium album Limber. The amino acid sequence of proteinase T as deduced from the nucleotide sequence is about 56% identical to that of proteinase K. |
| cd04059 | Peptidases_S8_Protein_convertases_Kexins_Furin-like | 9.83e-11 | 227 | 449 | 79 | 295 | Peptidase S8 family domain in Protein convertases. Protein convertases, whose members include furins and kexins, are members of the peptidase S8 or Subtilase clan of proteases. They have an Asp/His/Ser catalytic triad that is not homologous to trypsin. Kexins are involved in the activation of peptide hormones, growth factors, and viral proteins. Furin cleaves cell surface vasoactive peptides and proteins involved in cardiovascular tissue remodeling in the TGN, at cell surface, or in endosomes but rarely in the ER. Furin also plays a key role in blood pressure regulation though the activation of transforming growth factor (TGF)-beta. High specificity is seen for cleavage after dibasic (Lys-Arg or Arg-Arg) or multiple basic residues in protein convertases. There is also strong sequence conservation. |
| cd07473 | Peptidases_S8_Subtilisin_like | 1.00e-10 | 206 | 451 | 47 | 259 | Peptidase S8 family domain in Subtilisin-like proteins. This family is a member of the Peptidases S8 or Subtilases serine endo- and exo-peptidase clan. They have an Asp/His/Ser catalytic triad similar to that found in trypsin-like proteases, but do not share their three-dimensional structure and are not homologous to trypsin. The stability of subtilases may be enhanced by calcium, some members have been shown to bind up to 4 ions via binding sites with different affinity. Some members of this clan contain disulfide bonds. These enzymes can be intra- and extracellular, some function at extreme temperatures and pH values. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QLA69175.1 | 0.0 | 1 | 682 | 1 | 682 |
| QKZ97715.1 | 0.0 | 1 | 682 | 1 | 682 |
| ASQ19654.1 | 0.0 | 1 | 682 | 1 | 664 |
| QUG54103.1 | 0.0 | 1 | 682 | 1 | 664 |
| QGN44379.1 | 0.0 | 1 | 682 | 1 | 664 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
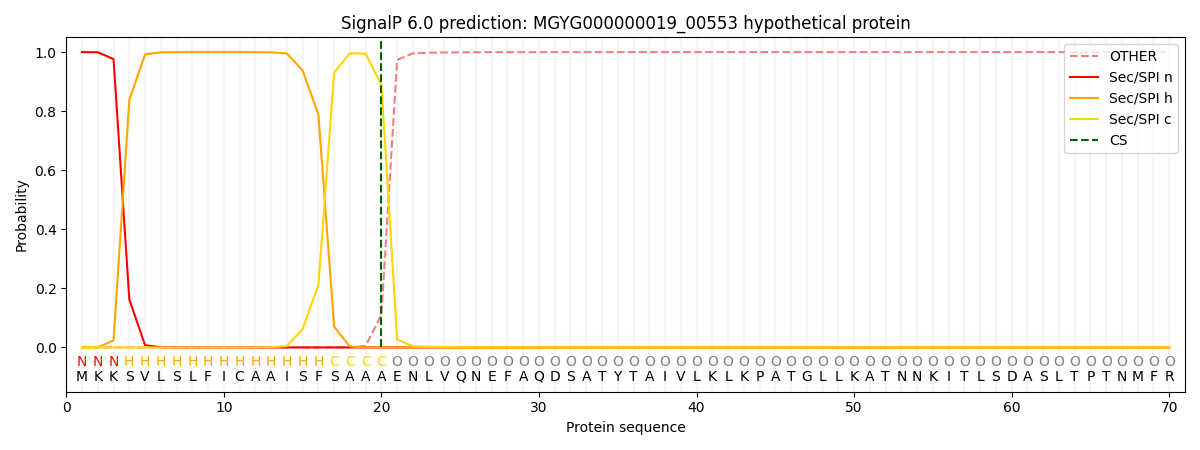
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000619 | 0.998341 | 0.000462 | 0.000191 | 0.000190 | 0.000184 |
