You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000021_01705
You are here: Home > Sequence: MGYG000000021_01705
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Limosilactobacillus vaginalis_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Limosilactobacillus; Limosilactobacillus vaginalis_A | |||||||||||
| CAZyme ID | MGYG000000021_01705 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 16098; End: 17846 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00877 | NLPC_P60 | 1.41e-14 | 283 | 389 | 1 | 102 | NlpC/P60 family. The function of this domain is unknown. It is found in several lipoproteins. |
| pfam06605 | Prophage_tail | 1.39e-13 | 409 | 534 | 125 | 251 | Prophage endopeptidase tail. This family is of prophage tail proteins that are probably acting as endopeptidases. |
| pfam18994 | Prophage_tailD1 | 8.43e-13 | 11 | 93 | 1 | 82 | Prophage endopeptidase tail N-terminal domain. This domain represents the N-terminal domain of prophage tail proteins that are probably acting as endopeptidases. This domain has a RIFT related fold. |
| NF033741 | NlpC_p60_RipA | 4.16e-10 | 235 | 380 | 274 | 442 | NlpC/P60 family peptidoglycan endopeptidase RipA. |
| pfam06605 | Prophage_tail | 1.67e-09 | 121 | 260 | 3 | 134 | Prophage endopeptidase tail. This family is of prophage tail proteins that are probably acting as endopeptidases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QYY85594.1 | 3.89e-111 | 10 | 565 | 3 | 657 |
| QHM66411.1 | 4.64e-109 | 10 | 565 | 3 | 657 |
| QHM64692.1 | 4.64e-109 | 10 | 565 | 3 | 657 |
| QHM69436.1 | 4.64e-109 | 10 | 565 | 3 | 657 |
| QDZ69499.1 | 1.41e-107 | 10 | 565 | 3 | 657 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6B8C_A | 5.83e-19 | 273 | 387 | 30 | 138 | Crystalstructure of NlpC/p60 domain of peptidoglycan hydrolase SagA [Enterococcus faecium] |
| 3NE0_A | 1.07e-11 | 259 | 380 | 61 | 199 | Structureand functional regulation of RipA, a mycobacterial enzyme essential for daughter cell separation [Mycobacterium tuberculosis H37Rv] |
| 3PBC_A | 1.07e-11 | 259 | 380 | 61 | 199 | ChainA, Invasion Protein [Mycobacterium tuberculosis] |
| 4Q4T_A | 7.87e-11 | 259 | 380 | 319 | 457 | Structureof the Resuscitation Promoting Factor Interacting protein RipA mutated at E444 [Mycobacterium tuberculosis H37Rv] |
| 3S0Q_A | 1.65e-10 | 259 | 380 | 62 | 200 | ChainA, INVASION PROTEIN [Mycobacterium tuberculosis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P13692 | 8.63e-16 | 273 | 387 | 404 | 512 | Protein P54 OS=Enterococcus faecium OX=1352 PE=3 SV=2 |
| O53168 | 4.31e-10 | 259 | 380 | 319 | 457 | Peptidoglycan endopeptidase RipA OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=ripA PE=1 SV=1 |
| A0QX22 | 9.89e-09 | 260 | 380 | 352 | 482 | Peptidoglycan endopeptidase RipA OS=Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155) OX=246196 GN=ripA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
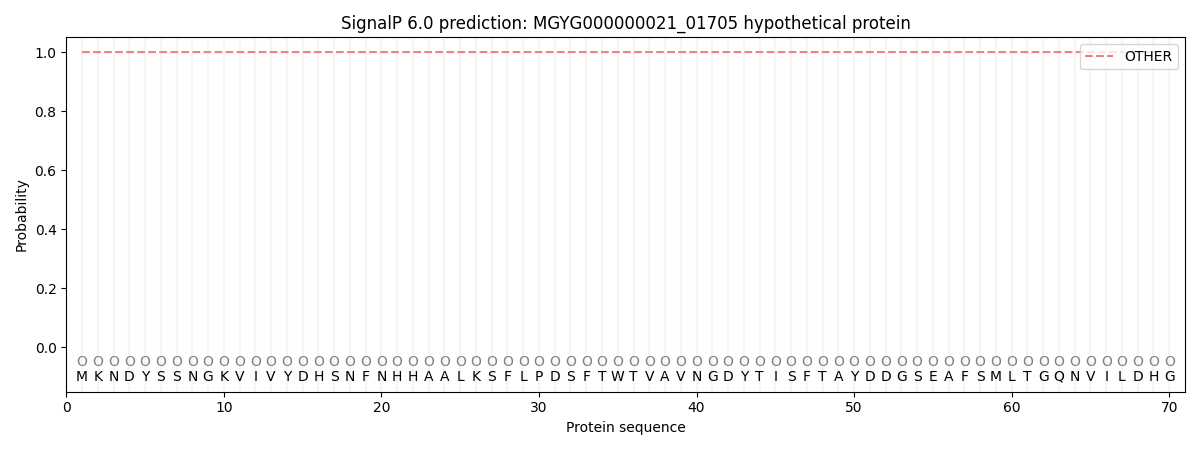
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000058 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
