You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000024_00851
You are here: Home > Sequence: MGYG000000024_00851
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
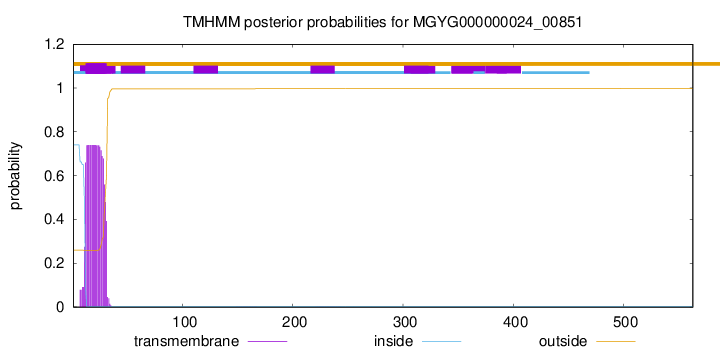
TMHMM annotations
Basic Information help
| Species | Paraclostridium bifermentans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Peptostreptococcales; Peptostreptococcaceae; Paraclostridium; Paraclostridium bifermentans | |||||||||||
| CAZyme ID | MGYG000000024_00851 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 883577; End: 885268 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06543 | GH18_PF-ChiA-like | 9.12e-92 | 63 | 374 | 1 | 294 | PF-ChiA is an uncharacterized chitinase found in the hyperthermophilic archaeon Pyrococcus furiosus with a glycosyl hydrolase family 18 (GH18) catalytic domain as well as a cellulose-binding domain. Members of this domain family are found not only in archaea but also in eukaryotes and prokaryotes. PF-ChiA exhibits hydrolytic activity toward both colloidal and crystalline (beta/alpha) chitins at high temperature. |
| pfam16403 | DUF5011 | 7.21e-16 | 392 | 463 | 2 | 71 | Domain of unknown function (DUF5011). This small family of proteins is functionally uncharacterized. This family is found in Bacteroides, Prevotella, and Parabateroides. Proteins in this family are around 230 amino acids in length. |
| cd12204 | CBD_like | 3.31e-10 | 532 | 562 | 17 | 47 | Cellulose-binding domain, chitinase and related proteins. This group contains proteins related to the cellulose-binding domain of Erwinia chrysanthemi endoglucanase Z (EGZ) and Serratia marcescens chitinase B (ChiB). Gram negative plant parasite Erwinia chrysanthemi produces a variety of depolymerizing enzymes to metabolize pectin and cellulose on the host plant. Cellulase EGZ has a modular structure, with N-terminal catalytic domain linked to a C-terminal cellulose-binding domain (CBD). CBD mediates the secretion activity of EGZ. Chitinases allow certain bacteria to utilize chitin as a energy source. Typically, non-plant chitinases are of the glycosidase family 18. |
| cd12215 | ChiC_BD | 1.70e-09 | 472 | 512 | 1 | 42 | Chitin-binding domain of chitinase C. Chitin-binding domain of chitinase C (ChiC) of Streptomyces griseus and related proteins. Chitinase C is a family 19 chitinase, and consists of a N-terminal chitin binding domain and a C-terminal chitin-catalytic domain that effects degradation. Chitinases function in invertebrates in the degradation of old exoskeletons, in fungi to utilize chitin in cell walls, and in bacteria which use chitin as an energy source. ChiC contains the characteristic chitin-binding aromatic residues. |
| cd02871 | GH18_chitinase_D-like | 6.56e-07 | 121 | 227 | 44 | 152 | GH18 domain of Chitinase D (ChiD). ChiD, a chitinase found in Bacillus circulans, hydrolyzes the 1,4-beta-linkages of N-acetylglucosamine in chitin and chitodextrins. The domain architecture of ChiD includes a catalytic glycosyl hydrolase family 18 (GH18) domain, a chitin-binding domain, and a fibronectin type III domain. The chitin-binding and fibronectin type III domains are located either N-terminal or C-terminal to the catalytic domain. This family includes exochitinase Chi36 from Bacillus cereus. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QEZ69311.1 | 0.0 | 1 | 563 | 1 | 563 |
| CDH91379.1 | 2.04e-256 | 2 | 562 | 5 | 585 |
| ACD22750.1 | 2.04e-256 | 2 | 562 | 5 | 585 |
| AIY82117.1 | 4.74e-255 | 11 | 562 | 17 | 585 |
| QDM47091.1 | 1.30e-227 | 52 | 563 | 691 | 1276 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2DSK_A | 2.68e-22 | 61 | 354 | 8 | 263 | Crystalstructure of catalytic domain of hyperthermophilic chitinase from Pyrococcus furiosus [Pyrococcus furiosus],2DSK_B Crystal structure of catalytic domain of hyperthermophilic chitinase from Pyrococcus furiosus [Pyrococcus furiosus] |
| 3A4W_A | 1.65e-21 | 61 | 354 | 8 | 263 | Crystalstructures of catalytic site mutants of active domain 2 of thermostable chitinase from Pyrococcus furiosus complexed with chito-oligosaccharides [Pyrococcus furiosus],3A4W_B Crystal structures of catalytic site mutants of active domain 2 of thermostable chitinase from Pyrococcus furiosus complexed with chito-oligosaccharides [Pyrococcus furiosus] |
| 3A4X_A | 3.01e-21 | 61 | 354 | 8 | 263 | Crystalstructures of catalytic site mutants of active domain 2 of thermostable chitinase from Pyrococcus furiosus complexed with chito-oligosaccharides [Pyrococcus furiosus],3A4X_B Crystal structures of catalytic site mutants of active domain 2 of thermostable chitinase from Pyrococcus furiosus complexed with chito-oligosaccharides [Pyrococcus furiosus],3AFB_A Crystal structures of catalytic site mutants of active domain 2 of chitinase from Pyrococcus furiosus [Pyrococcus furiosus],3AFB_B Crystal structures of catalytic site mutants of active domain 2 of chitinase from Pyrococcus furiosus [Pyrococcus furiosus] |
| 4HMC_A | 2.68e-16 | 281 | 510 | 299 | 524 | Crystalstructure of cold-adapted chitinase from Moritella marina [Moritella marina],4HMD_A Crystal structure of cold-adapted chitinase from Moritella marina with a reaction intermediate - oxazolinium ion (NGO) [Moritella marina],4HME_A Crystal structure of cold-adapted chitinase from Moritella marina with a reaction product - NAG2 [Moritella marina] |
| 4MB3_A | 2.68e-16 | 281 | 510 | 299 | 524 | Crystalstructure of E153Q mutant of cold-adapted chitinase from Moritella marina [Moritella marina],4MB4_A Crystal structure of E153Q mutant of cold-adapted chitinase from Moritella complex with Nag4 [Moritella marina],4MB5_A Crystal structure of E153Q mutant of cold-adapted chitinase from Moritella complex with Nag5 [Moritella marina] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P13656 | 2.24e-29 | 60 | 376 | 584 | 894 | Probable bifunctional chitinase/lysozyme OS=Escherichia coli (strain K12) OX=83333 GN=chiA PE=1 SV=2 |
| O74199 | 2.26e-27 | 60 | 354 | 233 | 509 | Endochitinase 11 OS=Metarhizium anisopliae OX=5530 GN=chi11 PE=1 SV=1 |
| P14529 | 3.79e-20 | 64 | 354 | 30 | 297 | Chitinase OS=Saccharopolyspora erythraea (strain ATCC 11635 / DSM 40517 / JCM 4748 / NBRC 13426 / NCIMB 8594 / NRRL 2338) OX=405948 GN=chiA2 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001440 | 0.993572 | 0.004108 | 0.000412 | 0.000225 | 0.000180 |