You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000029_00146
You are here: Home > Sequence: MGYG000000029_00146
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
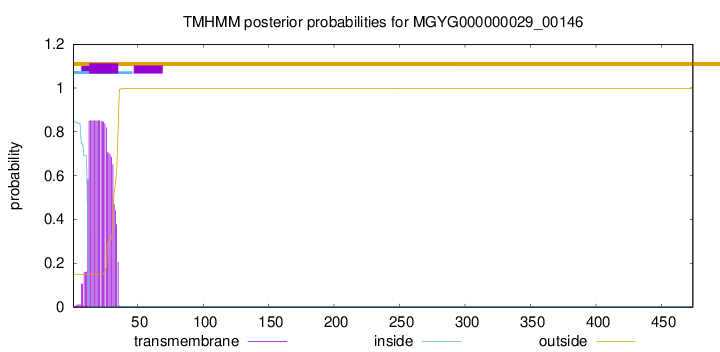
TMHMM annotations
Basic Information help
| Species | Bacteroides finegoldii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides finegoldii | |||||||||||
| CAZyme ID | MGYG000000029_00146 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 194169; End: 195593 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 200 | 455 | 1.1e-41 | 0.7476923076923077 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 4.17e-12 | 228 | 466 | 246 | 470 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam00295 | Glyco_hydro_28 | 1.11e-08 | 203 | 421 | 91 | 276 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN02793 | PLN02793 | 2.46e-05 | 229 | 386 | 186 | 330 | Probable polygalacturonase |
| PLN02218 | PLN02218 | 4.01e-04 | 228 | 464 | 200 | 423 | polygalacturonase ADPG |
| PLN03003 | PLN03003 | 0.001 | 230 | 423 | 148 | 332 | Probable polygalacturonase At3g15720 |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIU95271.1 | 0.0 | 26 | 474 | 19 | 467 |
| QRQ56356.1 | 2.39e-308 | 18 | 474 | 12 | 468 |
| ALJ44674.1 | 2.39e-308 | 18 | 474 | 12 | 468 |
| SCV09008.1 | 2.39e-308 | 18 | 474 | 12 | 468 |
| QNL37066.1 | 2.78e-307 | 18 | 474 | 12 | 468 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A2QW66 | 3.92e-06 | 191 | 431 | 162 | 389 | Probable exopolygalacturonase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=pgxA PE=3 SV=1 |
| Q27UB2 | 3.92e-06 | 191 | 431 | 162 | 389 | Exopolygalacturonase A OS=Aspergillus niger OX=5061 GN=pgxA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
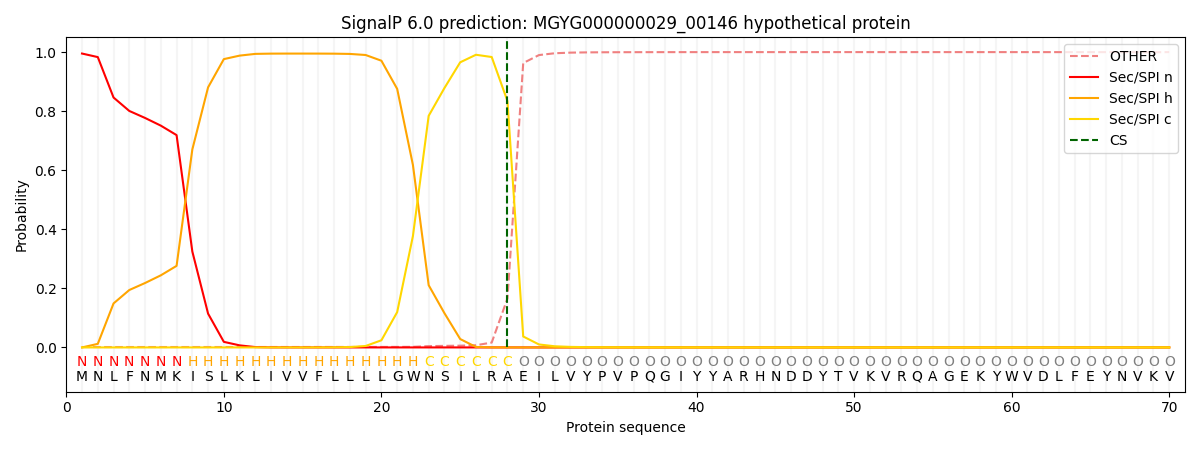
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.002618 | 0.992414 | 0.004269 | 0.000239 | 0.000224 | 0.000222 |