You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000029_02745
You are here: Home > Sequence: MGYG000000029_02745
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides finegoldii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides finegoldii | |||||||||||
| CAZyme ID | MGYG000000029_02745 | |||||||||||
| CAZy Family | GH26 | |||||||||||
| CAZyme Description | Mannan endo-1,4-beta-mannosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 13399; End: 14499 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH26 | 30 | 301 | 1e-62 | 0.8382838283828383 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02156 | Glyco_hydro_26 | 3.33e-33 | 31 | 306 | 6 | 284 | Glycosyl hydrolase family 26. |
| COG4124 | ManB2 | 3.70e-16 | 109 | 359 | 117 | 340 | Beta-mannanase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ47536.1 | 2.88e-274 | 1 | 366 | 1 | 366 |
| SCV06947.1 | 2.88e-274 | 1 | 366 | 1 | 366 |
| QRQ58988.1 | 2.88e-274 | 1 | 366 | 1 | 366 |
| QUR45660.1 | 4.10e-274 | 1 | 366 | 1 | 366 |
| QTO27343.1 | 4.41e-174 | 20 | 366 | 20 | 374 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4ZXO_A | 1.55e-260 | 23 | 366 | 1 | 344 | Thestructure of a GH26 beta-mannanase from Bacteroides ovatus, BoMan26A. [Bacteroides ovatus] |
| 2VX4_A | 2.39e-80 | 31 | 366 | 39 | 396 | CellvibrioJaponicus Mannanase Cjman26c Native Form [Cellvibrio japonicus],2VX6_A CELLVIBRIO JAPONICUS MANNANASE CJMAN26C Gal1Man4-BOUND FORM [Cellvibrio japonicus] |
| 4CD4_A | 4.61e-80 | 31 | 366 | 62 | 419 | Thestructure of GH26 beta-mannanase CjMan26C from Cellvibrio japonicus in complex with ManIFG [Cellvibrio japonicus Ueda107],4CD5_A The structure of GH26 beta-mannanase CjMan26C from Cellvibrio japonicus in complex with ManMIm [Cellvibrio japonicus Ueda107] |
| 2VX5_A | 4.77e-80 | 31 | 365 | 39 | 395 | CellvibrioJaponicus Mannanase Cjman26c Mannose-Bound Form [Cellvibrio japonicus] |
| 2VX7_A | 1.89e-79 | 31 | 366 | 39 | 396 | CellvibrioJaponicus Mannanase Cjman26c Mannobiose-Bound Form [Cellvibrio japonicus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P49424 | 5.61e-43 | 31 | 351 | 55 | 409 | Mannan endo-1,4-beta-mannosidase OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=manA PE=1 SV=2 |
| A1A278 | 5.32e-37 | 31 | 356 | 48 | 393 | Mannan endo-1,4-beta-mannosidase OS=Bifidobacterium adolescentis (strain ATCC 15703 / DSM 20083 / NCTC 11814 / E194a) OX=367928 GN=BAD_1030 PE=1 SV=1 |
| P16699 | 7.34e-22 | 12 | 325 | 15 | 330 | Mannan endo-1,4-beta-mannosidase A and B OS=Caldalkalibacillus mannanilyticus (strain DSM 16130 / CIP 109019 / JCM 10596 / AM-001) OX=1236954 PE=1 SV=1 |
| P55278 | 5.24e-19 | 35 | 323 | 45 | 322 | Mannan endo-1,4-beta-mannosidase OS=Bacillus subtilis OX=1423 PE=3 SV=1 |
| P49425 | 2.61e-18 | 72 | 305 | 189 | 424 | Mannan endo-1,4-beta-mannosidase OS=Rhodothermus marinus (strain ATCC 43812 / DSM 4252 / R-10) OX=518766 GN=manA PE=1 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
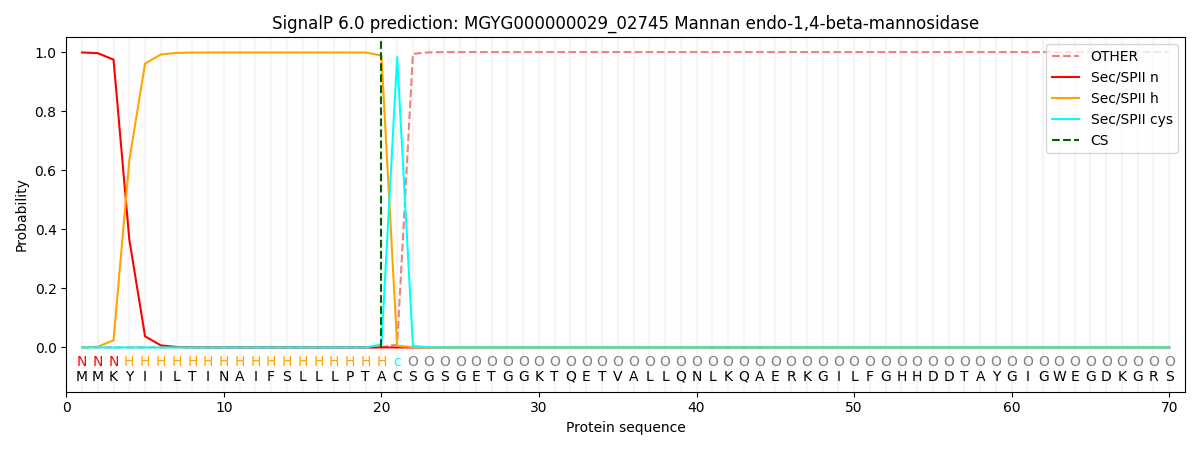
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000003 | 0.001248 | 0.998802 | 0.000001 | 0.000001 | 0.000001 |
