You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000041_01253
You are here: Home > Sequence: MGYG000000041_01253
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Acetatifactor sp900066365 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Acetatifactor; Acetatifactor sp900066365 | |||||||||||
| CAZyme ID | MGYG000000041_01253 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 61804; End: 64104 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00118 | LysM | 2.06e-13 | 649 | 694 | 2 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| pfam01476 | LysM | 2.52e-12 | 650 | 695 | 1 | 43 | LysM domain. The LysM (lysin motif) domain is about 40 residues long. It is found in a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. The structure of this domain is known. |
| pfam09479 | Flg_new | 3.66e-12 | 529 | 600 | 1 | 65 | Listeria-Bacteroides repeat domain (List_Bact_rpt). This model describes a conserved core region of about 43 residues, which occurs in at least two families of tandem repeats. These include 78-residue repeats which occur from 2 to 15 times in some proteins of Bacteroides forsythus ATCC 43037, and 70-residue repeats found in families of internalins of Listeria species. Single copies are found in proteins of Fibrobacter succinogenes, Geobacter sulfurreducens, and a few other bacteria. |
| smart00257 | LysM | 9.89e-12 | 649 | 694 | 1 | 44 | Lysin motif. |
| NF033188 | internalin_H | 1.72e-11 | 400 | 632 | 294 | 513 | InlH/InlC2 family class 1 internalin. Internalins, as found in the intracellular human pathogen Listeria monocytogenes, are paralogous surface or secreted proteins with an N-terminal signal peptide, leucine-rich repeats, and usually a C-terminal LPXTG processing and cell surface anchoring site. See PMID:17764999 for a general discussion of internalins. Members of this family are internalin H (InlH), or internalin C2, two class 1 (LPXTG-type) internalins that are closely related, one apparently derived from the other through a recombination event. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNM02905.1 | 5.51e-267 | 1 | 766 | 1 | 723 |
| QNM02904.1 | 3.66e-210 | 1 | 766 | 1 | 864 |
| QIB56830.1 | 2.90e-14 | 637 | 766 | 235 | 350 |
| QBE96304.1 | 2.90e-14 | 637 | 766 | 235 | 350 |
| QMW80393.1 | 2.90e-14 | 637 | 766 | 235 | 350 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4B8V_A | 5.19e-06 | 645 | 764 | 39 | 161 | ChainA, Extracellular Protein 6 [Fulvia fulva],4B9H_A Chain A, Extracellular Protein 6 [Fulvia fulva] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0DJ97 | 2.00e-15 | 444 | 601 | 655 | 800 | Putative Gly-rich membrane protein Bcell_0380 OS=Evansella cellulosilytica (strain ATCC 21833 / DSM 2522 / FERM P-1141 / JCM 9156 / N-4) OX=649639 GN=Bcell_0380 PE=4 SV=1 |
| P0DJ98 | 7.68e-14 | 444 | 630 | 75 | 238 | Putative membrane protein Bcell_0381 OS=Evansella cellulosilytica (strain ATCC 21833 / DSM 2522 / FERM P-1141 / JCM 9156 / N-4) OX=649639 GN=Bcell_0381 PE=4 SV=1 |
| O31852 | 2.83e-08 | 649 | 763 | 28 | 131 | D-gamma-glutamyl-meso-diaminopimelic acid endopeptidase CwlS OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlS PE=1 SV=1 |
| O07532 | 3.41e-08 | 649 | 763 | 175 | 283 | Peptidoglycan endopeptidase LytF OS=Bacillus subtilis (strain 168) OX=224308 GN=lytF PE=1 SV=2 |
| Q37896 | 4.91e-07 | 649 | 763 | 164 | 262 | Endolysin OS=Bacillus phage B103 OX=10778 GN=15 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
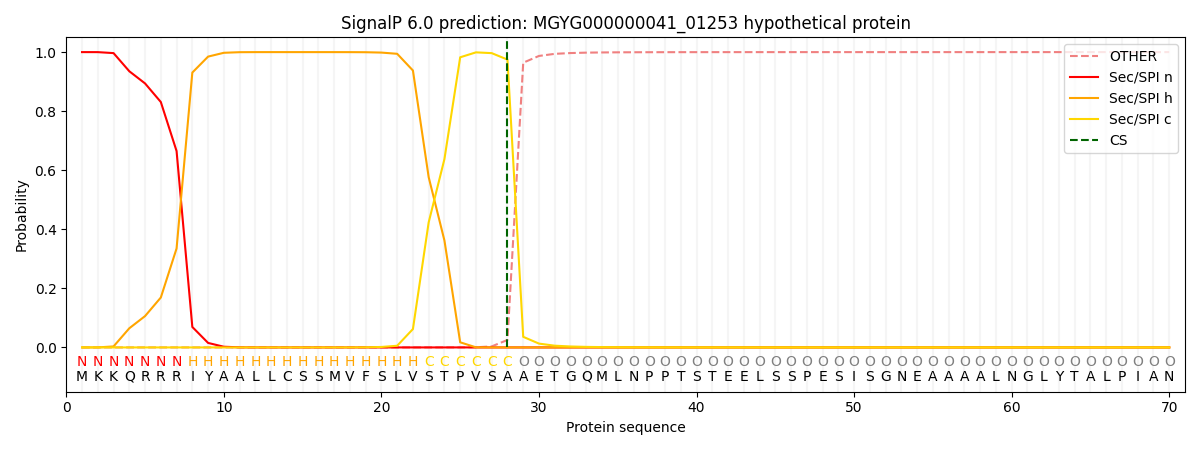
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000289 | 0.998972 | 0.000179 | 0.000206 | 0.000177 | 0.000145 |