You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000054_01779
You are here: Home > Sequence: MGYG000000054_01779
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
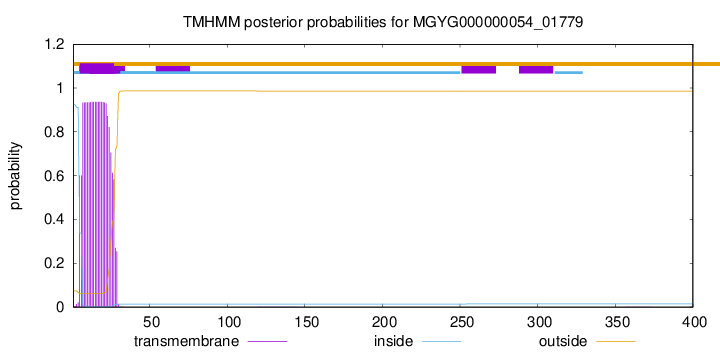
TMHMM annotations
Basic Information help
| Species | Bacteroides acidifaciens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides acidifaciens | |||||||||||
| CAZyme ID | MGYG000000054_01779 | |||||||||||
| CAZy Family | PL38 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 32878; End: 34080 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL38 | 66 | 344 | 2.2e-97 | 0.996415770609319 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam05426 | Alginate_lyase | 3.19e-96 | 65 | 344 | 1 | 274 | Alginate lyase. This family contains several bacterial alginate lyase proteins. Alginate is a family of 1-4-linked copolymers of beta -D-mannuronic acid (M) and alpha -L-guluronic acid (G). It is produced by brown algae and by some bacteria belonging to the genera Azotobacter and Pseudomonas. Alginate lyases catalyze the depolymerization of alginates by beta -elimination, generating a molecule containing 4-deoxy-L-erythro-hex-4-enepyranosyluronate at the nonreducing end. This family adopts an all alpha fold. |
| PRK00325 | algL | 2.30e-05 | 237 | 308 | 194 | 270 | polysaccharide lyase. |
| cd00244 | AlgLyase | 0.002 | 237 | 353 | 178 | 290 | Alginate Lyase A1-III; enzymatically depolymerizes alginate, a complex copolymer of beta-D-mannuronate and alpha-L-guluronate, by cleaving the beta-(1,4) glycosidic bond. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT26274.1 | 5.63e-296 | 1 | 400 | 1 | 400 |
| QIU93437.1 | 5.63e-296 | 1 | 400 | 1 | 400 |
| QNL39956.1 | 4.08e-289 | 1 | 400 | 1 | 400 |
| QNT66841.1 | 5.92e-120 | 24 | 382 | 23 | 380 |
| QEC55803.1 | 5.63e-79 | 26 | 380 | 34 | 387 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3NFV_A | 3.76e-65 | 55 | 385 | 37 | 380 | Crystalstructure of an alginate lyase (BACOVA_01668) from Bacteroides ovatus at 1.95 A resolution [Bacteroides ovatus ATCC 8483],3NNB_A Crystal structure of an alginate lyase (BACOVA_01668) from Bacteroides ovatus at 1.60 A resolution [Bacteroides ovatus ATCC 8483] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
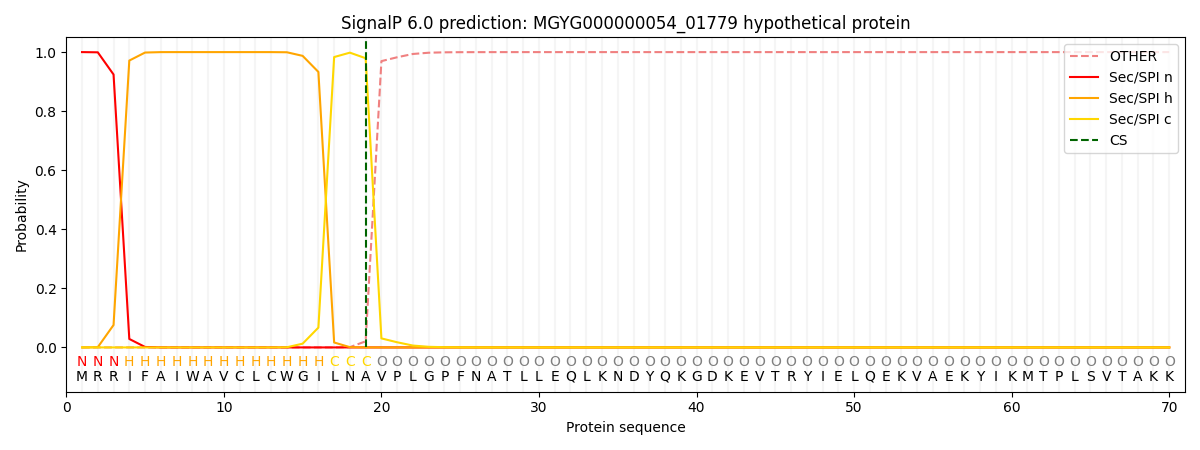
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000219 | 0.999212 | 0.000155 | 0.000135 | 0.000136 | 0.000128 |