You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000054_03002
You are here: Home > Sequence: MGYG000000054_03002
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides acidifaciens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides acidifaciens | |||||||||||
| CAZyme ID | MGYG000000054_03002 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 35500; End: 36810 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 73 | 414 | 1.1e-59 | 0.9261538461538461 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.07e-54 | 49 | 365 | 85 | 417 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam00295 | Glyco_hydro_28 | 1.02e-14 | 132 | 345 | 43 | 242 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN03003 | PLN03003 | 6.13e-11 | 142 | 289 | 112 | 259 | Probable polygalacturonase At3g15720 |
| PLN02218 | PLN02218 | 2.98e-10 | 49 | 265 | 70 | 272 | polygalacturonase ADPG |
| PLN03010 | PLN03010 | 9.23e-09 | 44 | 317 | 44 | 280 | polygalacturonase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ43485.1 | 2.32e-234 | 1 | 431 | 1 | 431 |
| BCA48691.1 | 4.68e-234 | 1 | 431 | 1 | 431 |
| QQA10748.1 | 2.70e-233 | 1 | 431 | 1 | 431 |
| QUT42011.1 | 7.72e-233 | 1 | 431 | 1 | 431 |
| QUT69827.1 | 7.72e-233 | 1 | 431 | 1 | 431 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OLP_A | 3.73e-36 | 45 | 434 | 43 | 451 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
| 3JUR_A | 4.59e-26 | 45 | 321 | 26 | 328 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 4MXN_A | 3.28e-19 | 39 | 287 | 14 | 235 | Crystalstructure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_B Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_C Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_D Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184] |
| 2UVE_A | 1.77e-11 | 49 | 202 | 159 | 310 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
| 4C2L_A | 9.50e-06 | 78 | 263 | 45 | 216 | Crystalstructure of endo-xylogalacturonan hydrolase from Aspergillus tubingensis [Aspergillus tubingensis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7PZL3 | 1.31e-20 | 49 | 265 | 65 | 279 | Probable polygalacturonase OS=Vitis vinifera OX=29760 GN=GSVIVT00026920001 PE=1 SV=1 |
| Q949Z1 | 8.32e-15 | 23 | 320 | 56 | 331 | Polygalacturonase At1g48100 OS=Arabidopsis thaliana OX=3702 GN=At1g48100 PE=2 SV=1 |
| P43212 | 5.09e-13 | 47 | 284 | 59 | 282 | Polygalacturonase OS=Cryptomeria japonica OX=3369 PE=1 SV=1 |
| Q9LW07 | 7.54e-11 | 142 | 318 | 112 | 291 | Probable polygalacturonase At3g15720 OS=Arabidopsis thaliana OX=3702 GN=At3g15720 PE=3 SV=1 |
| P27644 | 3.04e-09 | 183 | 301 | 23 | 138 | Polygalacturonase OS=Rhizobium radiobacter OX=358 GN=pgl PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
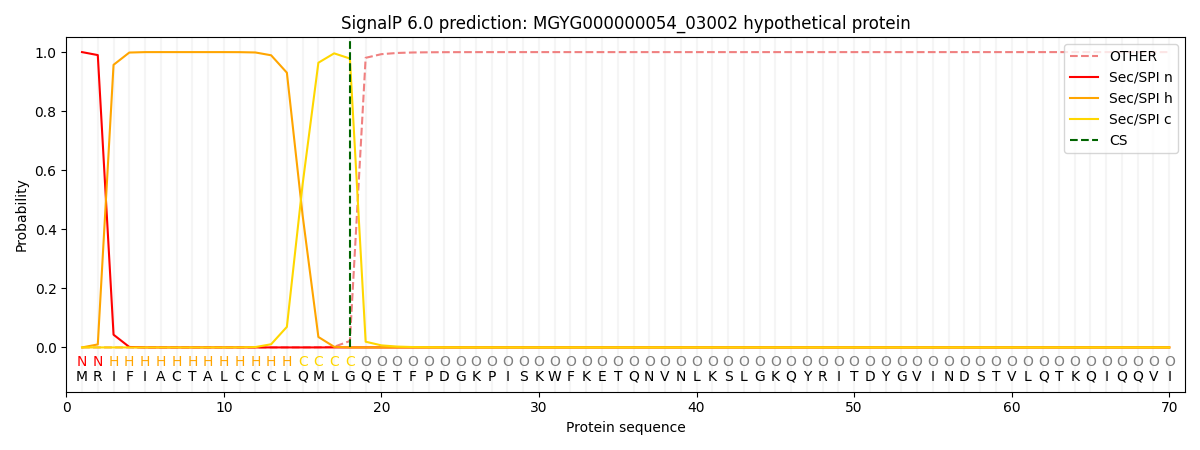
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000262 | 0.999104 | 0.000206 | 0.000138 | 0.000134 | 0.000127 |
