You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000054_04408
You are here: Home > Sequence: MGYG000000054_04408
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides acidifaciens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides acidifaciens | |||||||||||
| CAZyme ID | MGYG000000054_04408 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4736; End: 5608 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 34 | 283 | 3.3e-53 | 0.973568281938326 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0627 | FrmB | 5.80e-27 | 42 | 289 | 36 | 313 | S-formylglutathione hydrolase FrmB [Defense mechanisms]. |
| pfam00756 | Esterase | 5.13e-24 | 34 | 277 | 1 | 239 | Putative esterase. This family contains Esterase D. However it is not clear if all members of the family have the same function. This family is related to the pfam00135 family. |
| COG2382 | Fes | 2.32e-22 | 30 | 277 | 71 | 288 | Enterochelin esterase or related enzyme [Inorganic ion transport and metabolism]. |
| COG4947 | COG4947 | 9.12e-07 | 123 | 277 | 89 | 223 | Esterase/lipase superfamily enzyme [General function prediction only]. |
| COG0657 | Aes | 1.76e-06 | 43 | 270 | 66 | 287 | Acetyl esterase/lipase [Lipid transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| SNV45150.1 | 1.47e-27 | 34 | 286 | 152 | 392 |
| QNA46493.1 | 2.39e-27 | 24 | 286 | 40 | 277 |
| AVM58192.1 | 7.76e-27 | 33 | 282 | 154 | 394 |
| VTQ01161.1 | 8.05e-27 | 34 | 277 | 42 | 273 |
| QUT65236.1 | 5.41e-26 | 33 | 282 | 154 | 394 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5VOL_A | 1.19e-157 | 19 | 289 | 4 | 273 | Bacint_04212ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_B Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_C Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_D Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_E Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_F Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_G Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393],5VOL_H Bacint_04212 ferulic acid esterase [Bacteroides intestinalis DSM 17393] |
| 6RZN_A | 1.75e-22 | 34 | 286 | 139 | 386 | Crystalstructure of the N-terminal carbohydrate binding module family 48 and ferulic acid esterase from the multi-enzyme CE1-GH62-GH10 [uncultured bacterium],6RZN_B Crystal structure of the N-terminal carbohydrate binding module family 48 and ferulic acid esterase from the multi-enzyme CE1-GH62-GH10 [uncultured bacterium] |
| 7B5V_A | 6.95e-20 | 33 | 158 | 145 | 275 | ChainA, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B5V_B Chain B, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B6B_A Chain A, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836],7B6B_B Chain B, Carbohydrate Esterase family 1 protein with an N-terminal carbohydrate binding module family 48 [Dysgonomonas mossii DSM 22836] |
| 1JJF_A | 1.97e-17 | 24 | 278 | 29 | 251 | ChainA, Endo-1,4-beta-xylanase Z [Acetivibrio thermocellus] |
| 6MOT_A | 3.90e-17 | 33 | 158 | 123 | 252 | ChainA, Isoamylase N-terminal domain protein [Bacteroides intestinalis DSM 17393] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EXZ4 | 3.07e-17 | 33 | 158 | 437 | 566 | Carbohydrate acetyl esterase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axe1-6A PE=1 SV=1 |
| P10478 | 6.61e-16 | 24 | 278 | 48 | 270 | Endo-1,4-beta-xylanase Z OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xynZ PE=1 SV=3 |
| P31471 | 4.05e-15 | 31 | 219 | 147 | 346 | Uncharacterized protein YieL OS=Escherichia coli (strain K12) OX=83333 GN=yieL PE=4 SV=3 |
| D5EY13 | 6.21e-12 | 31 | 277 | 504 | 717 | Endo-1,4-beta-xylanase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyn10D-fae1A PE=1 SV=1 |
| P23553 | 1.75e-08 | 27 | 286 | 3 | 263 | Acetyl esterase OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=xynC PE=4 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
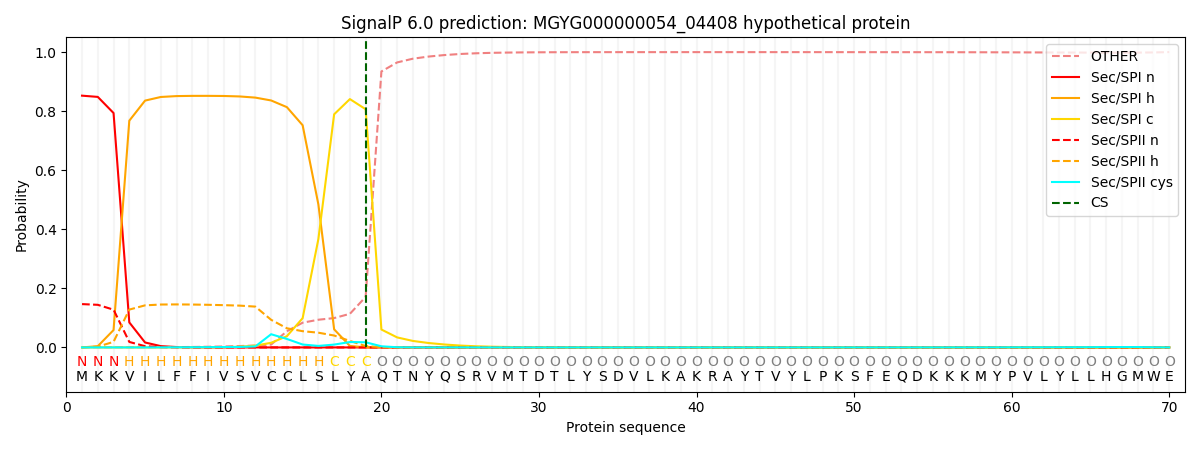
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.002159 | 0.843540 | 0.153302 | 0.000355 | 0.000309 | 0.000299 |
