You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000055_00714
You are here: Home > Sequence: MGYG000000055_00714
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Muribaculum intestinale | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; Muribaculum; Muribaculum intestinale | |||||||||||
| CAZyme ID | MGYG000000055_00714 | |||||||||||
| CAZy Family | GH53 | |||||||||||
| CAZyme Description | Arabinogalactan endo-beta-1,4-galactanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 120853; End: 121908 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH53 | 44 | 327 | 3.4e-92 | 0.9005847953216374 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07745 | Glyco_hydro_53 | 3.03e-97 | 43 | 348 | 1 | 333 | Glycosyl hydrolase family 53. This domain belongs to family 53 of the glycosyl hydrolase classification. These enzymes are enzymes are endo-1,4- beta-galactanases (EC:3.2.1.89). The structure of this domain is known and has a TIM barrel fold. |
| COG3867 | GanB | 2.80e-83 | 42 | 331 | 39 | 353 | Arabinogalactan endo-1,4-beta-galactosidase [Carbohydrate transport and metabolism]. |
| cd03311 | CIMS_C_terminal_like | 0.009 | 144 | 236 | 145 | 237 | CIMS - Cobalamine-independent methonine synthase, or MetE, C-terminal domain_like. Many members have been characterized as 5-methyltetrahydropteroyltriglutamate-homocysteine methyltransferases, EC:2.1.1.14, mostly from bacteria and plants. This enzyme catalyses the last step in the production of methionine by transferring a methyl group from 5-methyltetrahydrofolate to L-homocysteine without using an intermediate methyl carrier. The active enzyme has a dual (beta-alpha)8-barrel structure, and this model covers the C-terminal barrel, and a few single-barrel sequences most similar to the C-terminal barrel. It is assumed that the homologous N-terminal barrel has evolved from the C-terminus via gene duplication and has subsequently lost binding sites, and it seems as if the two barrels forming the active enzyme may sometimes reside on different polypeptides. The C-terminal domain incorporates the Zinc ion, which binds and activates homocysteine. Sidechains from both barrels contribute to the binding of the folate substrate. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QMW86976.1 | 1.57e-223 | 1 | 351 | 3 | 353 |
| AAO79773.1 | 1.57e-223 | 1 | 351 | 3 | 353 |
| QUT42647.1 | 9.06e-223 | 1 | 351 | 3 | 353 |
| BCA49296.1 | 9.06e-223 | 1 | 351 | 3 | 353 |
| QUT70301.1 | 1.29e-222 | 1 | 351 | 3 | 353 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6GP5_A | 4.14e-220 | 17 | 351 | 18 | 352 | Beta-1,4-galactanasefrom Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],6GP5_B Beta-1,4-galactanase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
| 6GPA_A | 6.95e-212 | 36 | 349 | 1 | 314 | Beta-1,4-galactanasefrom Bacteroides thetaiotaomicron with galactose [Bacteroides thetaiotaomicron VPI-5482],6GPA_B Beta-1,4-galactanase from Bacteroides thetaiotaomicron with galactose [Bacteroides thetaiotaomicron VPI-5482] |
| 1HJS_A | 4.38e-41 | 44 | 338 | 5 | 313 | Structureof two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_B Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_C Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJS_D Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_A Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_B Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_C Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus],1HJU_D Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and pH optimum. [Thermothelomyces thermophilus] |
| 7OSK_A | 1.10e-39 | 42 | 290 | 50 | 292 | ChainA, Arabinogalactan endo-1,4-beta-galactosidase [Ignisphaera aggregans DSM 17230],7OSK_B Chain B, Arabinogalactan endo-1,4-beta-galactosidase [Ignisphaera aggregans DSM 17230] |
| 4BF7_A | 1.39e-39 | 44 | 338 | 22 | 330 | Emericillanidulans endo-beta-1,4-galactanase [Aspergillus nidulans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P48843 | 1.59e-49 | 38 | 295 | 2 | 254 | Uncharacterized protein in bgaB 5'region (Fragment) OS=Niallia circulans OX=1397 PE=3 SV=1 |
| P83692 | 2.40e-40 | 44 | 338 | 5 | 313 | Arabinogalactan endo-beta-1,4-galactanase OS=Thermothelomyces thermophilus OX=78579 PE=1 SV=1 |
| Q5B153 | 7.29e-39 | 44 | 338 | 22 | 330 | Arabinogalactan endo-beta-1,4-galactanase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=galA PE=1 SV=2 |
| A1D3T4 | 1.61e-38 | 44 | 351 | 26 | 349 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=galA PE=3 SV=1 |
| B0XPR3 | 4.39e-38 | 44 | 338 | 26 | 336 | Probable arabinogalactan endo-beta-1,4-galactanase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=galA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
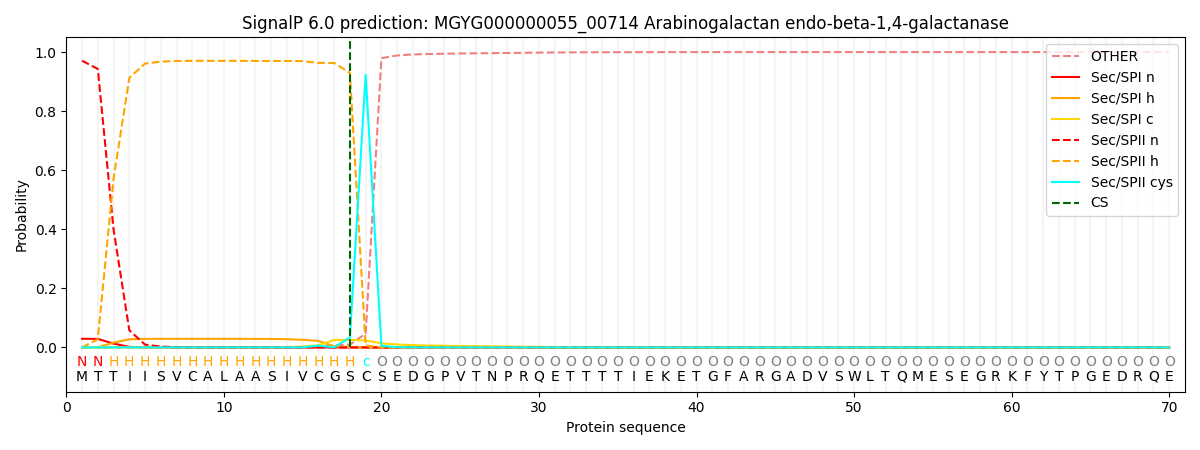
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000091 | 0.028476 | 0.971417 | 0.000011 | 0.000015 | 0.000012 |
