You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000057_00297
You are here: Home > Sequence: MGYG000000057_00297
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp002491635 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp002491635 | |||||||||||
| CAZyme ID | MGYG000000057_00297 | |||||||||||
| CAZy Family | GH76 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 72423; End: 73556 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH76 | 55 | 352 | 1.1e-86 | 0.8379888268156425 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03663 | Glyco_hydro_76 | 9.78e-53 | 35 | 332 | 2 | 315 | Glycosyl hydrolase family 76. Family of alpha-1,6-mannanases. |
| COG4833 | COG4833 | 5.49e-48 | 35 | 346 | 3 | 323 | Predicted alpha-1,6-mannanase, GH76 family [Carbohydrate transport and metabolism]. |
| cd04434 | LanC_like | 4.75e-05 | 117 | 346 | 1 | 216 | Cyclases involved in the biosynthesis of lantibiotics, and similar proteins. LanC is the cyclase enzyme of the lanthionine synthetase. Lanthionine is a lantibiotic, a unique class of peptide antibiotics. They are ribosomally synthesized as a precursor peptide and then post-translationally modified to contain thioether cross-links called lanthionines (Lans) or methyllanthionines (MeLans), in addition to 2,3-didehydroalanine (Dha) and (Z)-2,3-didehydrobutyrine (Dhb). These unusual amino acids are introduced by the dehydration of serine and threonine residues, followed by thioether formation via addition of cysteine thiols, catalysed by LanB and LanC or LanM. LanC, the cyclase component, is a zinc metalloprotein, whose bound metal has been proposed to activate the thiol substrate for nucleophilic addition. A related domain is also present in LanM and other pro- and eukaryotic proteins with poorly characterized functions. |
| cd04792 | LanM-like | 0.003 | 123 | 267 | 499 | 622 | Cyclases involved in the biosynthesis of class II lantibiotics, and similar proteins. LanM-like proteins. LanM is a bifunctional enzyme, involved in the synthesis of class II lantibiotics. It is responsible for both the dehydration and the cyclization of the precursor-peptide during lantibiotic synthesis. The C-terminal domain shows similarity to LanC, the cyclase component of the lan operon, but the N terminus seems to be unrelated to the dehydratase, LanB. |
| pfam05147 | LANC_like | 0.009 | 123 | 274 | 14 | 142 | Lanthionine synthetase C-like protein. Lanthionines are thioether bridges that are putatively generated by dehydration of Ser and Thr residues followed by addition of cysteine residues within the peptide. This family contains the lanthionine synthetase C-like proteins 1 and 2 which are related to the bacterial lanthionine synthetase components C (LanC). LANCL1 (P40 seven-transmembrane-domain protein) and LANCL2 (testes-specific adriamycin sensitivity protein) are thought to be peptide-modifying enzyme components in eukaryotic cells. Both proteins are produced in large quantities in the brain and testes and may have role in the immune surveillance of these organs. Lanthionines are found in lantibiotics, which are peptide-derived, post-translationally modified antimicrobials produced by several bacterial strains. This region contains seven internal repeats. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUU01224.1 | 1.84e-262 | 1 | 377 | 1 | 377 |
| QUT37354.1 | 1.84e-262 | 1 | 377 | 1 | 377 |
| QQA30005.1 | 3.72e-262 | 1 | 377 | 1 | 377 |
| BBK86488.1 | 3.72e-262 | 1 | 377 | 1 | 377 |
| QUT65780.1 | 3.72e-262 | 1 | 377 | 1 | 377 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6SHD_A | 1.03e-86 | 34 | 373 | 53 | 390 | Structureof the GH76A alpha-1,6-mannanase from Salegentibacter sp. HEL1_6 [Salegentibacter sp. Hel_I_6],6SHD_B Structure of the GH76A alpha-1,6-mannanase from Salegentibacter sp. HEL1_6 [Salegentibacter sp. Hel_I_6],6SHD_C Structure of the GH76A alpha-1,6-mannanase from Salegentibacter sp. HEL1_6 [Salegentibacter sp. Hel_I_6] |
| 6Y8F_A | 3.34e-85 | 34 | 373 | 54 | 391 | ChainA, Alpha-1,6-endo-mannanase GH76A mutant [Salegentibacter sp. Hel_I_6] |
| 6SHM_A | 2.66e-84 | 34 | 373 | 54 | 391 | Aninactive (D136A and D137A) variant of alpha-1,6-mannanase, GH76A of Salegentibacter sp. HEL1_6 in complex with alpha-1,6-mannotetrose [Salegentibacter sp. Hel_I_6] |
| 3K7X_A | 5.28e-55 | 67 | 373 | 41 | 341 | ChainA, Lin0763 protein [Listeria innocua] |
| 4D4A_A | 6.24e-38 | 23 | 331 | 15 | 325 | ChainA, Alpha-1,6-mannanase [Niallia circulans],4D4A_B Chain B, Alpha-1,6-mannanase [Niallia circulans],4D4B_A Chain A, Alpha-1,6-mannanase [Niallia circulans],4D4B_B Chain B, Alpha-1,6-mannanase [Niallia circulans],4D4C_A Chain A, Alpha-1,6-mannanase [Niallia circulans],4D4C_B Chain B, Alpha-1,6-mannanase [Niallia circulans],4D4D_A Chain A, Alpha-1,6-mannanase [Niallia circulans],4D4D_B Chain B, Alpha-1,6-mannanase [Niallia circulans],5N0F_A Chain A, Alpha-1,6-mannanase [Niallia circulans],5N0F_B Chain B, Alpha-1,6-mannanase [Niallia circulans],6ZBX_A Chain A, Alpha-1,6-mannanase [Niallia circulans],6ZBX_B Chain B, Alpha-1,6-mannanase [Niallia circulans],7NL5_A Chain A, Alpha-1,6-mannanase [Niallia circulans] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as LIPO
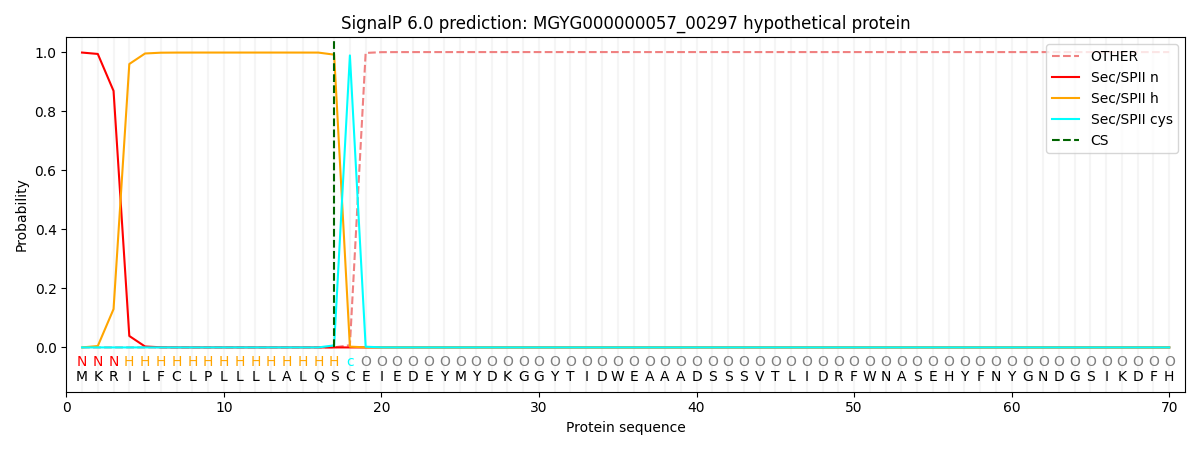
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000002 | 0.001503 | 0.998560 | 0.000000 | 0.000001 | 0.000001 |
