You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000057_03297
You are here: Home > Sequence: MGYG000000057_03297
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
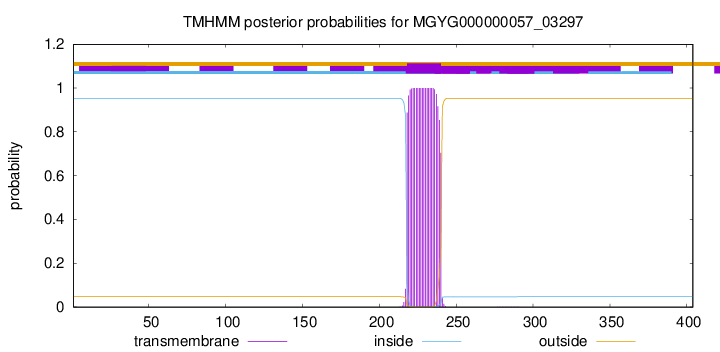
TMHMM annotations
Basic Information help
| Species | Bacteroides sp002491635 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp002491635 | |||||||||||
| CAZyme ID | MGYG000000057_03297 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | Integration host factor subunit alpha | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4416; End: 5630 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00216 | Bac_DNA_binding | 9.39e-21 | 6 | 93 | 1 | 88 | Bacterial DNA-binding protein. |
| COG0776 | HimA | 3.04e-17 | 4 | 96 | 1 | 93 | Bacterial nucleoid DNA-binding protein [Replication, recombination and repair]. |
| cd13835 | IHF_A | 1.70e-16 | 8 | 93 | 3 | 88 | Alpha subunit of integration host factor (IHFA). This subfamily consists of the alpha subunit of integration host factor (IHF) and IHF-like domains. IHF is a nucleoid-associated protein (NAP) that binds and sharply bends many DNA targets in a sequence specific manner. It is a heterodimeric protein composed of two highly homologous subunits IHFA (IHF-alpha) and IHFB (IHF-beta). It is known to act as a transcription factor at many gene regulatory regions in E. coli. IHF is an essential cofactor in phage lambda site-specific recombination, having an architectural role during assembly of specialized nucleoprotein structures (snups). IHF is also involved in formation as well as maintenance of bacterial biofilms since it is found in complex with extracellular DNA (eDNA) within the extracellular polymeric substances (EPS) matrix of many biofilms. |
| cd00591 | HU_IHF | 4.31e-16 | 8 | 91 | 2 | 85 | DNA sequence specific (IHF) and non-specific (HU) domains. This family includes integration host factor (IHF) and HU, also called type II DNA-binding proteins (DNABII), which are small dimeric proteins that specifically bind the DNA minor groove, inducing large bends in the DNA and serving as architectural factors in a variety of cellular processes such as recombination, initiation of replication/transcription and gene regulation. IHF binds DNA in a sequence specific manner while HU displays little or no sequence preference. IHF homologs are usually heterodimers, while HU homologs are typically homodimers (except HU heterodimers from E. coli and other enterobacteria). HU is highly basic and contributes to chromosomal compaction and maintenance of negative supercoiling, thus often referred to as histone-like protein. IHF is an essential cofactor in phage lambda site-specific recombination, having an architectural role during assembly of specialized nucleoprotein structures (snups). Bacillus phage SPO1-encoded transcription factor 1 (TF1) is another related type II DNA-binding protein. Like IHF, TF1 binds DNA specifically and bends DNA sharply. |
| cd13832 | IHF | 6.17e-16 | 9 | 91 | 3 | 85 | Integration host factor (IHF) and similar proteins. This subfamily includes integration host factor (IHF) and IHF-like domains. IHF is a nucleoid-associated protein (NAP) that binds and sharply bends many DNA targets in a sequence specific manner. It is a heterodimeric protein composed of two highly homologous subunits IHFA (IHF-alpha) and IHFB (IHF-beta). It is known to act as a transcription factor at many gene regulatory regions in E. coli. IHF is an essential cofactor in phage lambda site-specific recombination, having an architectural role during assembly of specialized nucleoprotein structures (snups). IHF is also involved in formation as well as maintenance of bacterial biofilms since it is found in complex with extracellular DNA (eDNA) within the extracellular polymeric substances (EPS) matrix of many biofilms. This subfamily also includes the protein Hbb from tick-borne spirochete Borrelia burgdorferi, responsible for causing Lyme disease in humans. Hbb, a homodimer, shows DNA sequence preferences that are related, yet distinct from those of IHF. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QPH57227.1 | 4.36e-212 | 1 | 404 | 1 | 393 |
| QBJ16968.1 | 4.36e-212 | 1 | 404 | 1 | 393 |
| QMI81676.1 | 4.36e-212 | 1 | 404 | 1 | 393 |
| QQA31617.1 | 9.71e-209 | 1 | 404 | 1 | 393 |
| BBK88465.1 | 9.71e-209 | 1 | 404 | 1 | 393 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1HUE_A | 3.49e-11 | 5 | 93 | 1 | 89 | ChainA, HU PROTEIN [Geobacillus stearothermophilus],1HUE_B Chain B, HU PROTEIN [Geobacillus stearothermophilus],1HUU_A Dna-Binding Protein Hu From Bacillus Stearothermophilus [Geobacillus stearothermophilus],1HUU_B Dna-Binding Protein Hu From Bacillus Stearothermophilus [Geobacillus stearothermophilus],1HUU_C Dna-Binding Protein Hu From Bacillus Stearothermophilus [Geobacillus stearothermophilus] |
| 5LVT_A | 4.36e-10 | 6 | 93 | 3 | 90 | Structureof HU protein from Lactococcus lactis [Lactococcus lactis subsp. lactis Il1403],5LVT_B Structure of HU protein from Lactococcus lactis [Lactococcus lactis subsp. lactis Il1403],5LVT_C Structure of HU protein from Lactococcus lactis [Lactococcus lactis subsp. lactis Il1403],5LVT_D Structure of HU protein from Lactococcus lactis [Lactococcus lactis subsp. lactis Il1403] |
| 4QJN_A | 2.48e-09 | 5 | 93 | 1 | 89 | ChainA, DNA-binding protein HU [Staphylococcus aureus subsp. aureus Mu50],4QJN_B Chain B, DNA-binding protein HU [Staphylococcus aureus subsp. aureus Mu50],4QJN_C Chain C, DNA-binding protein HU [Staphylococcus aureus subsp. aureus Mu50],4QJN_D Chain D, DNA-binding protein HU [Staphylococcus aureus subsp. aureus Mu50],4QJU_A Chain A, DNA-binding protein HU [Staphylococcus aureus subsp. aureus Mu50],4QJU_B Chain B, DNA-binding protein HU [Staphylococcus aureus subsp. aureus Mu50] |
| 5J0N_I | 6.01e-09 | 9 | 93 | 6 | 90 | ChainI, Integration host factor subunit alpha [Escherichia coli],5J0N_K Chain K, Integration host factor subunit alpha [Escherichia coli] |
| 1IHF_A | 6.47e-09 | 9 | 93 | 7 | 91 | INTEGRATIONHOST FACTOR/DNA COMPLEX [Escherichia coli],1OUZ_A Chain A, Integration Host Factor Alpha-subunit [Escherichia coli],1OWF_A Chain A, Integration Host Factor Alpha-subunit [Escherichia coli],1OWG_A Crystal structure of WT IHF complexed with an altered H' site (T44A) [Escherichia coli],2HT0_A IHF bound to doubly nicked DNA [Escherichia coli],5WFE_K Cas1-Cas2-IHF-DNA holo-complex [Escherichia coli S88] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B8J836 | 1.50e-11 | 9 | 107 | 5 | 103 | Integration host factor subunit alpha OS=Anaeromyxobacter dehalogenans (strain 2CP-1 / ATCC BAA-258) OX=455488 GN=ihfA PE=3 SV=1 |
| B4UAP1 | 1.50e-11 | 9 | 107 | 5 | 103 | Integration host factor subunit alpha OS=Anaeromyxobacter sp. (strain K) OX=447217 GN=ihfA PE=3 SV=1 |
| Q2IJA7 | 2.05e-11 | 9 | 94 | 5 | 90 | Integration host factor subunit alpha OS=Anaeromyxobacter dehalogenans (strain 2CP-C) OX=290397 GN=ihfA PE=3 SV=1 |
| A7HBJ3 | 5.03e-11 | 9 | 94 | 5 | 90 | Integration host factor subunit alpha OS=Anaeromyxobacter sp. (strain Fw109-5) OX=404589 GN=ihfA PE=3 SV=1 |
| Q65TL4 | 6.71e-11 | 9 | 95 | 7 | 93 | Integration host factor subunit alpha OS=Mannheimia succiniciproducens (strain MBEL55E) OX=221988 GN=ihfA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000032 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |