You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000067_02086
You are here: Home > Sequence: MGYG000000067_02086
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
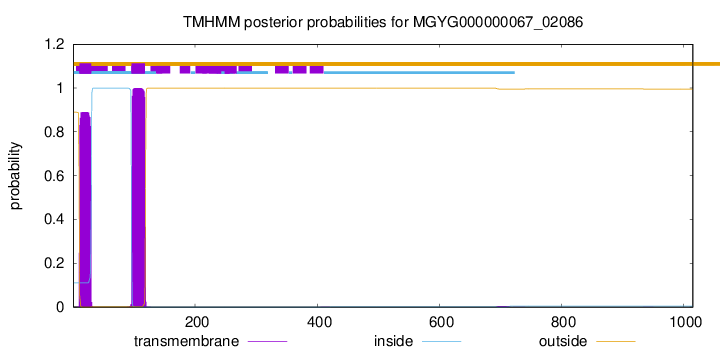
TMHMM annotations
Basic Information help
| Species | Lachnoclostridium_B sp900066765 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Lachnoclostridium_B; Lachnoclostridium_B sp900066765 | |||||||||||
| CAZyme ID | MGYG000000067_02086 | |||||||||||
| CAZy Family | GT4 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 95153; End: 98200 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd03794 | GT4_WbuB-like | 3.21e-92 | 620 | 1007 | 1 | 388 | Escherichia coli WbuB and similar proteins. This family is most closely related to the GT1 family of glycosyltransferases. WbuB in E. coli is involved in the biosynthesis of the O26 O-antigen. It has been proposed to function as an N-acetyl-L-fucosamine (L-FucNAc) transferase. |
| pfam02397 | Bac_transf | 1.79e-72 | 91 | 270 | 1 | 181 | Bacterial sugar transferase. This Pfam family represents a conserved region from a number of different bacterial sugar transferases, involved in diverse biosynthesis pathways. |
| COG2148 | WcaJ | 1.06e-69 | 86 | 275 | 36 | 226 | Sugar transferase involved in LPS biosynthesis (colanic, teichoic acid) [Cell wall/membrane/envelope biogenesis]. |
| cd05232 | UDP_G4E_4_SDR_e | 3.34e-67 | 312 | 603 | 1 | 303 | UDP-glucose 4 epimerase, subgroup 4, extended (e) SDRs. UDP-glucose 4 epimerase (aka UDP-galactose-4-epimerase), is a homodimeric extended SDR. It catalyzes the NAD-dependent conversion of UDP-galactose to UDP-glucose, the final step in Leloir galactose synthesis. This subgroup is comprised of bacterial proteins, and includes the Staphylococcus aureus capsular polysaccharide Cap5N, which may have a role in the synthesis of UDP-N-acetyl-d-fucosamine. This subgroup has the characteristic active site tetrad and NAD-binding motif of the extended SDRs. Extended SDRs are distinct from classical SDRs. In addition to the Rossmann fold (alpha/beta folding pattern with a central beta-sheet) core region typical of all SDRs, extended SDRs have a less conserved C-terminal extension of approximately 100 amino acids. Extended SDRs are a diverse collection of proteins, and include isomerases, epimerases, oxidoreductases, and lyases; they typically have a TGXXGXXG cofactor binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold, an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Sequence identity between different SDR enzymes is typically in the 15-30% range; they catalyze a wide range of activities including the metabolism of steroids, cofactors, carbohydrates, lipids, aromatic compounds, and amino acids, and act in redox sensing. Classical SDRs have an TGXXX[AG]XG cofactor binding motif and a YXXXK active site motif, with the Tyr residue of the active site motif serving as a critical catalytic residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase numbering). In addition to the Tyr and Lys, there is often an upstream Ser and/or an Asn, contributing to the active site; while substrate binding is in the C-terminal region, which determines specificity. The standard reaction mechanism is a 4-pro-S hydride transfer and proton relay involving the conserved Tyr and Lys, a water molecule stabilized by Asn, and nicotinamide. Atypical SDRs generally lack the catalytic residues characteristic of the SDRs, and their glycine-rich NAD(P)-binding motif is often different from the forms normally seen in classical or extended SDRs. Complex (multidomain) SDRs such as ketoreductase domains of fatty acid synthase have a GGXGXXG NAD(P)-binding motif and an altered active site motif (YXXXN). Fungal type ketoacyl reductases have a TGXXXGX(1-2)G NAD(P)-binding motif. |
| TIGR03025 | EPS_sugtrans | 7.93e-56 | 87 | 273 | 254 | 443 | exopolysaccharide biosynthesis polyprenyl glycosylphosphotransferase. Members of this family are generally found near other genes involved in the biosynthesis of a variety of exopolysaccharides. These proteins consist of two fused domains, an N-terminal hydrophobic domain of generally low conservation and a highly conserved C-terminal sugar transferase domain (pfam02397). Characterized and partially characterized members of this subfamily include Salmonella WbaP (originally RfbP), E. coli WcaJ, Methylobacillus EpsB, Xanthomonas GumD, Vibrio CpsA, Erwinia AmsG, Group B Streptococcus CpsE (originally CpsD), and Streptococcus suis Cps2E. Each of these is believed to act in transferring the sugar from, for instance, UDP-glucose or UDP-galactose, to a lipid carrier such as undecaprenyl phosphate as the first (priming) step in the synthesis of an oligosaccharide "block". This function is encoded in the C-terminal domain. The liposaccharide is believed to be subsequently transferred through a "flippase" function from the cytoplasmic to the periplasmic face of the inner membrane by the N-terminal domain. Certain closely related transferase enzymes, such as Sinorhizobium ExoY and Lactococcus EpsD, lack the N-terminal domain and are not found by this model. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUO31262.1 | 4.89e-274 | 1 | 652 | 1 | 664 |
| QMW80818.1 | 4.83e-189 | 90 | 633 | 34 | 654 |
| QIB56408.1 | 4.83e-189 | 90 | 633 | 34 | 654 |
| QEI31620.1 | 9.58e-184 | 617 | 1014 | 3 | 396 |
| QHB24119.1 | 9.58e-184 | 617 | 1014 | 3 | 396 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5W7L_A | 1.43e-29 | 85 | 284 | 5 | 200 | Structureof Campylobacter concisus PglC I57M/Q175M variant [Campylobacter concisus 13826],5W7L_B Structure of Campylobacter concisus PglC I57M/Q175M variant [Campylobacter concisus 13826] |
| 1HZJ_A | 5.01e-06 | 302 | 545 | 29 | 291 | HumanUdp-Galactose 4-Epimerase: Accommodation Of Udp-N- Acetylglucosamine Within The Active Site [Homo sapiens],1HZJ_B Human Udp-Galactose 4-Epimerase: Accommodation Of Udp-N- Acetylglucosamine Within The Active Site [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0P9D0 | 1.27e-28 | 85 | 278 | 1 | 189 | Undecaprenyl phosphate N,N'-diacetylbacillosamine 1-phosphate transferase OS=Campylobacter jejuni subsp. jejuni serotype O:2 (strain ATCC 700819 / NCTC 11168) OX=192222 GN=pglC PE=1 SV=1 |
| P71241 | 1.21e-27 | 87 | 266 | 270 | 455 | UDP-glucose:undecaprenyl-phosphate glucose-1-phosphate transferase OS=Escherichia coli (strain K12) OX=83333 GN=wcaJ PE=1 SV=2 |
| P10498 | 2.76e-27 | 87 | 269 | 2 | 194 | Exopolysaccharide production protein PSS OS=Rhizobium leguminosarum bv. phaseoli OX=385 GN=pss PE=3 SV=1 |
| Q44576 | 8.09e-25 | 86 | 265 | 333 | 522 | Undecaprenyl-phosphate glucose phosphotransferase OS=Komagataeibacter xylinus OX=28448 GN=aceA PE=3 SV=1 |
| Q57491 | 4.66e-24 | 78 | 273 | 262 | 469 | Uncharacterized sugar transferase HI_0872 OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=HI_0872 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999621 | 0.000376 | 0.000012 | 0.000002 | 0.000001 | 0.000005 |