You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000075_03083
You are here: Home > Sequence: MGYG000000075_03083
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
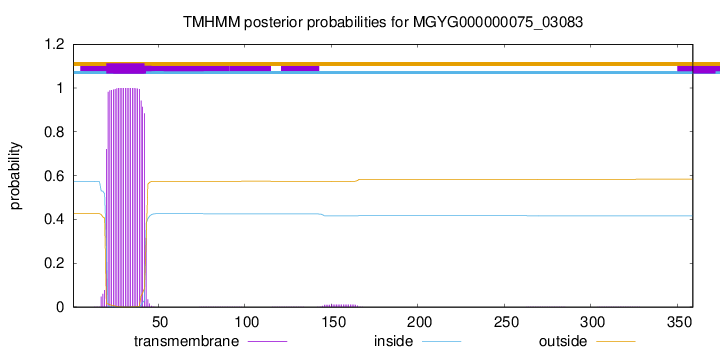
TMHMM annotations
Basic Information help
| Species | Paraclostridium tenue | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Peptostreptococcales; Peptostreptococcaceae; Paraclostridium; Paraclostridium tenue | |||||||||||
| CAZyme ID | MGYG000000075_03083 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 6752; End: 7831 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16891 | CwlT-like | 7.25e-61 | 68 | 209 | 1 | 150 | CwlT-like N-terminal lysozyme domain and similar domains. CwlT is a bifunctional cell wall hydrolase containing an N-terminal lysozyme domain and a C-terminal NlpC/P60 endopeptidase domain (gamma-d-D-glutamyl-L-diamino acid endopeptidase), and has been implicated in the spread of transposons. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| pfam13702 | Lysozyme_like | 9.34e-52 | 62 | 209 | 1 | 165 | Lysozyme-like. |
| pfam00877 | NLPC_P60 | 3.52e-28 | 261 | 355 | 14 | 105 | NlpC/P60 family. The function of this domain is unknown. It is found in several lipoproteins. |
| COG0791 | Spr | 1.43e-20 | 176 | 353 | 20 | 196 | Cell wall-associated hydrolase, NlpC family [Cell wall/membrane/envelope biogenesis]. |
| PRK10838 | spr | 1.44e-17 | 262 | 351 | 93 | 179 | bifunctional murein DD-endopeptidase/murein LD-carboxypeptidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QEZ70805.1 | 3.62e-164 | 1 | 357 | 1 | 363 |
| AWX74641.1 | 3.59e-141 | 57 | 359 | 66 | 368 |
| AVI31007.1 | 3.59e-141 | 57 | 359 | 66 | 368 |
| QTG87454.1 | 3.59e-141 | 57 | 359 | 66 | 368 |
| API45191.1 | 5.09e-141 | 57 | 359 | 66 | 368 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4HPE_A | 5.60e-33 | 50 | 348 | 9 | 298 | ChainA, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_B Chain B, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_C Chain C, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_D Chain D, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_E Chain E, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630],4HPE_F Chain F, Putative cell wall hydrolase Tn916-like,CTn1-Orf17 [Clostridioides difficile 630] |
| 4FDY_A | 1.21e-30 | 59 | 347 | 20 | 301 | ChainA, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50],4FDY_B Chain B, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50] |
| 2K1G_A | 3.82e-12 | 262 | 348 | 32 | 115 | SolutionNMR structure of lipoprotein spr from Escherichia coli K12. Northeast Structural Genomics target ER541-37-162 [Escherichia coli K-12] |
| 3PBI_A | 1.49e-07 | 238 | 335 | 101 | 192 | ChainA, Invasion Protein [Mycobacterium tuberculosis] |
| 3NE0_A | 1.21e-06 | 261 | 332 | 123 | 189 | Structureand functional regulation of RipA, a mycobacterial enzyme essential for daughter cell separation [Mycobacterium tuberculosis H37Rv] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P96645 | 5.19e-51 | 52 | 347 | 45 | 319 | Probable endopeptidase YddH OS=Bacillus subtilis (strain 168) OX=224308 GN=yddH PE=3 SV=1 |
| Q65NQ9 | 5.82e-43 | 223 | 359 | 318 | 452 | Peptidoglycan DL-endopeptidase CwlO OS=Bacillus licheniformis (strain ATCC 14580 / DSM 13 / JCM 2505 / CCUG 7422 / NBRC 12200 / NCIMB 9375 / NCTC 10341 / NRRL NRS-1264 / Gibson 46) OX=279010 GN=cwlO PE=3 SV=1 |
| P40767 | 2.26e-42 | 229 | 359 | 343 | 473 | Peptidoglycan DL-endopeptidase CwlO OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlO PE=1 SV=2 |
| O34636 | 3.07e-19 | 68 | 205 | 55 | 200 | Uncharacterized membrane protein YocA OS=Bacillus subtilis (strain 168) OX=224308 GN=yocA PE=4 SV=1 |
| P45296 | 7.21e-13 | 224 | 350 | 62 | 176 | Probable endopeptidase NlpC homolog OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=nlpC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
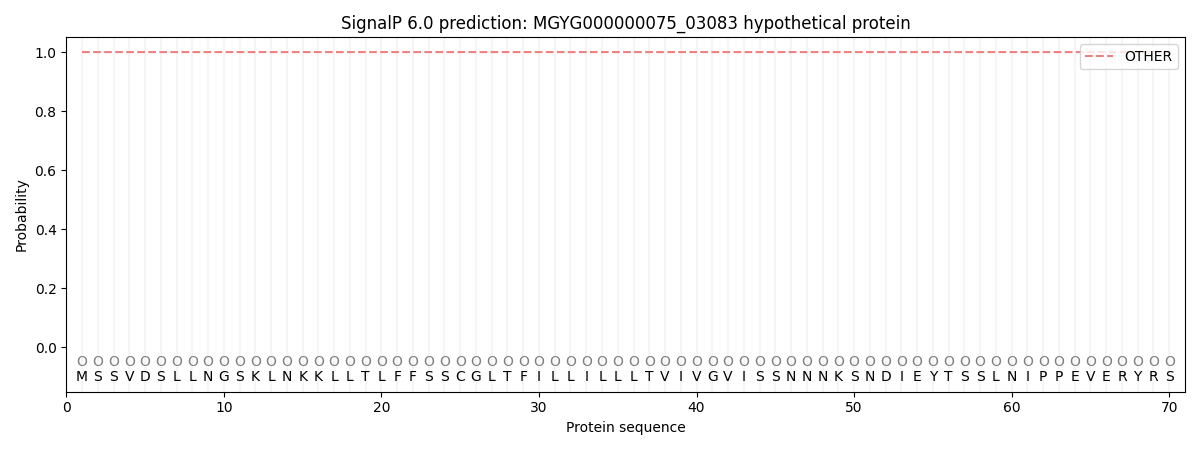
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000039 | 0.000004 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |