You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000082_02809
You are here: Home > Sequence: MGYG000000082_02809
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Acinetobacter mesopotamicus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Pseudomonadales; Moraxellaceae; Acinetobacter; Acinetobacter mesopotamicus | |||||||||||
| CAZyme ID | MGYG000000082_02809 | |||||||||||
| CAZy Family | GH153 | |||||||||||
| CAZyme Description | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 18317; End: 19558 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH153 | 65 | 405 | 2.2e-119 | 0.9914040114613181 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam14883 | GHL13 | 5.60e-132 | 69 | 386 | 1 | 325 | Hypothetical glycosyl hydrolase family 13. GHL13 is a family of hypothetical glycoside hydrolases. |
| TIGR03938 | deacetyl_PgaB | 1.31e-128 | 65 | 406 | 271 | 618 | poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB. Two well-characterized systems produce polysaccharide based on N-acetyl-D-glucosamine in straight chains with beta-1,6 linkages. These are encoded by the icaADBC operon in Staphylococcus species, where the system is designated polysaccharide intercellular adhesin (PIA), and the pgaABCD operon in Gram-negative bacteria such as E. coli. Both systems include a putative polysaccharide deacetylase. The PgaB protein, described here, contains an additional domain lacking from its Gram-positive counterpart IcaB (TIGR03933). Deacetylation by this protein appears necessary to allow export through the porin PgaA [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides] |
| PRK14582 | pgaB | 1.91e-98 | 31 | 405 | 286 | 663 | poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB. |
| PRK14581 | hmsF | 1.19e-84 | 33 | 405 | 288 | 664 | outer membrane N-deacetylase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AUC08208.1 | 3.90e-304 | 5 | 413 | 1 | 409 |
| QPF34269.1 | 3.90e-304 | 5 | 413 | 1 | 409 |
| QPF37286.1 | 3.90e-304 | 5 | 413 | 1 | 409 |
| QKQ69542.1 | 4.65e-242 | 1 | 413 | 1 | 413 |
| AYA02585.1 | 1.47e-185 | 11 | 405 | 3 | 404 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4P7L_A | 3.97e-78 | 66 | 405 | 10 | 359 | Structureof Escherichia coli PgaB C-terminal domain, P212121 crystal form [Escherichia coli K-12],4P7N_A Structure of Escherichia coli PgaB C-terminal domain in complex with glucosamine [Escherichia coli K-12],4P7O_A Structure of Escherichia coli PgaB C-terminal domain, P1 crystal form [Escherichia coli K-12],4P7O_B Structure of Escherichia coli PgaB C-terminal domain, P1 crystal form [Escherichia coli K-12],4P7Q_A Structure of Escherichia coli PgaB C-terminal domain in complex with N-acetylglucosamine [Escherichia coli K-12],4P7R_A Structure of Escherichia coli PgaB C-terminal domain in complex with a poly-beta-1,6-N-acetyl-D-glucosamine (PNAG) hexamer [Escherichia coli K-12] |
| 4F9D_A | 6.92e-74 | 66 | 395 | 278 | 617 | Structureof Escherichia coli PgaB 42-655 in complex with nickel [Escherichia coli K-12],4F9D_B Structure of Escherichia coli PgaB 42-655 in complex with nickel [Escherichia coli K-12] |
| 6AU1_A | 3.08e-71 | 66 | 406 | 8 | 355 | Structureof the PgaB (BpsB) glycoside hydrolase domain from Bordetella bronchiseptica [Bordetella bronchiseptica RB50],6AU1_B Structure of the PgaB (BpsB) glycoside hydrolase domain from Bordetella bronchiseptica [Bordetella bronchiseptica RB50] |
| 4F9J_A | 1.19e-69 | 66 | 395 | 278 | 617 | Structureof Escherichia coli PgaB 42-655 in complex with iron [Escherichia coli K-12],4F9J_B Structure of Escherichia coli PgaB 42-655 in complex with iron [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P75906 | 3.91e-74 | 66 | 405 | 315 | 664 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Escherichia coli (strain K12) OX=83333 GN=pgaB PE=1 SV=1 |
| Q8XAR3 | 3.91e-74 | 66 | 405 | 315 | 664 | Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase OS=Escherichia coli O157:H7 OX=83334 GN=pgaB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
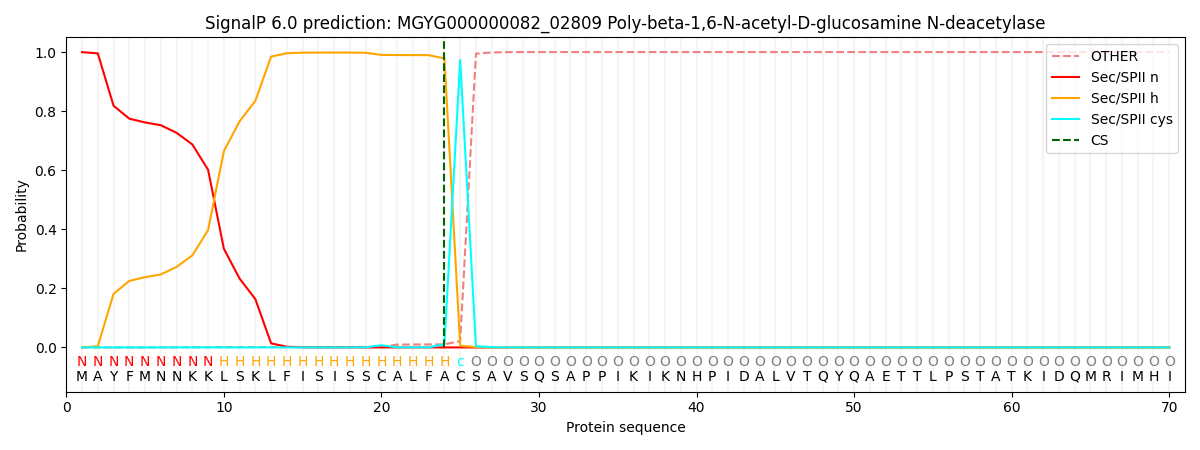
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000003 | 0.000060 | 0.999983 | 0.000001 | 0.000000 | 0.000000 |
