You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000083_02775
You are here: Home > Sequence: MGYG000000083_02775
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Peribacillus simplex | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales_B; DSM-1321; Peribacillus; Peribacillus simplex | |||||||||||
| CAZyme ID | MGYG000000083_02775 | |||||||||||
| CAZy Family | GH39 | |||||||||||
| CAZyme Description | HTH-type transcriptional activator RhaS | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 445545; End: 447863 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH39 | 333 | 718 | 7e-22 | 0.9559164733178654 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3664 | XynB | 1.33e-50 | 343 | 771 | 1 | 427 | Beta-xylosidase [Carbohydrate transport and metabolism]. |
| 225117 | COG2207 | 9.36e-25 | 165 | 265 | 19 | 119 | AraC AraC-type DNA-binding domain and AraC-containing proteins [Transcription]. |
| smart00342 | HTH_ARAC | 2.77e-22 | 184 | 265 | 3 | 84 | helix_turn_helix, arabinose operon control protein. |
| pfam12833 | HTH_18 | 9.72e-19 | 188 | 265 | 1 | 79 | Helix-turn-helix domain. |
| COG4977 | GlxA | 1.14e-15 | 165 | 265 | 219 | 319 | Transcriptional regulator GlxA family, contains an amidase domain and an AraC-type DNA-binding HTH domain [Transcription]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QYF80459.1 | 0.0 | 1 | 772 | 1 | 772 |
| QHS24385.1 | 4.54e-150 | 27 | 769 | 4 | 756 |
| QOF64550.1 | 7.88e-139 | 10 | 769 | 7 | 777 |
| QOF56795.1 | 7.88e-139 | 10 | 769 | 7 | 777 |
| QOF62367.1 | 2.39e-137 | 4 | 769 | 2 | 777 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6SWI_A | 6.82e-07 | 168 | 265 | 7 | 105 | TheC-terminal domain of AraT, a response regulator from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q6GKK1 | 7.19e-68 | 27 | 769 | 22 | 740 | Uncharacterized HTH-type transcriptional regulator SAR0107 OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=SAR0107 PE=4 SV=1 |
| Q5HJR8 | 1.36e-67 | 27 | 769 | 22 | 740 | Uncharacterized HTH-type transcriptional regulator SACOL0084 OS=Staphylococcus aureus (strain COL) OX=93062 GN=SACOL0084 PE=4 SV=2 |
| Q99XB1 | 6.76e-67 | 27 | 769 | 22 | 740 | Uncharacterized HTH-type transcriptional regulator SAV0101 OS=Staphylococcus aureus (strain Mu50 / ATCC 700699) OX=158878 GN=SAV0101 PE=4 SV=1 |
| Q7A882 | 6.76e-67 | 27 | 769 | 22 | 740 | Uncharacterized HTH-type transcriptional regulator SA0097 OS=Staphylococcus aureus (strain N315) OX=158879 GN=SA0097 PE=4 SV=1 |
| Q8NYT6 | 1.28e-66 | 27 | 769 | 22 | 740 | Uncharacterized HTH-type transcriptional regulator MW0077 OS=Staphylococcus aureus (strain MW2) OX=196620 GN=MW0077 PE=4 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
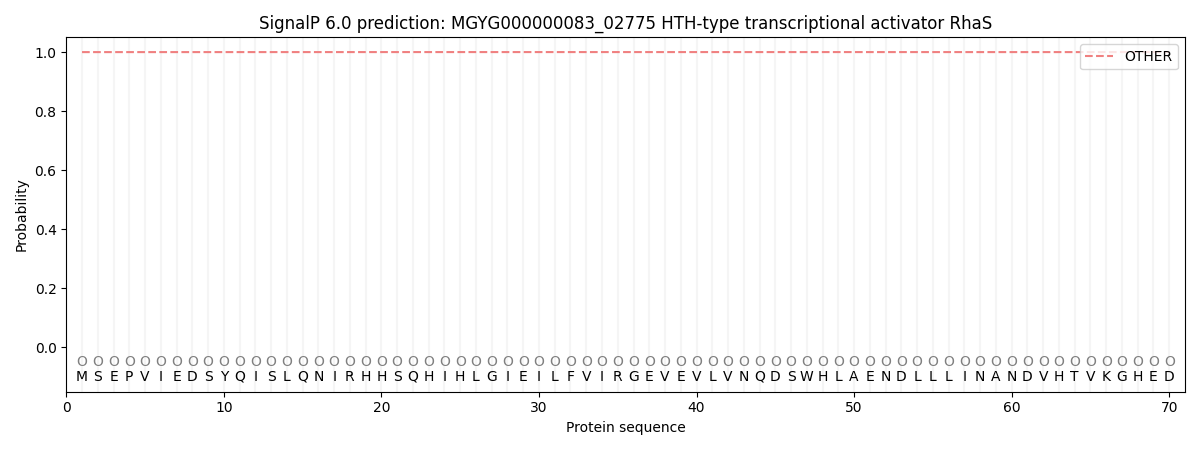
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000051 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
