You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000091_03646
You are here: Home > Sequence: MGYG000000091_03646
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Achromobacter xylosoxidans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Burkholderiales; Burkholderiaceae; Achromobacter; Achromobacter xylosoxidans | |||||||||||
| CAZyme ID | MGYG000000091_03646 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | D-alanine--poly(phosphoribitol) ligase subunit 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 657142; End: 669273 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3321 | PksD | 0.0 | 1549 | 2648 | 3 | 1059 | Acyl transferase domain in polyketide synthase (PKS) enzymes [Secondary metabolites biosynthesis, transport and catabolism]. |
| cd00833 | PKS | 0.0 | 1550 | 1974 | 1 | 421 | polyketide synthases (PKSs) polymerize simple fatty acids into a large variety of different products, called polyketides, by successive decarboxylating Claisen condensations. PKSs can be divided into 2 groups, modular type I PKSs consisting of one or more large multifunctional proteins and iterative type II PKSs, complexes of several monofunctional subunits. |
| cd12116 | A_NRPS_Ta1_like | 2.89e-170 | 582 | 1013 | 3 | 437 | The adenylation domain of nonribosomal peptide synthetases (NRPS), including salinosporamide A polyketide synthase. The adenylation (A) domain of NRPS recognizes a specific amino acid or hydroxy acid and activates it as an (amino) acyl adenylate by hydrolysis of ATP. The activated acyl moiety then forms a thioester to the enzyme-bound cofactor phosphopantetheine of a peptidyl carrier protein domain. NRPSs are large multifunctional enzymes which synthesize many therapeutically useful peptides in bacteria and fungi via a template-directed, nucleic acid independent nonribosomal mechanism. These natural products include antibiotics, immunosuppressants, plant and animal toxins, and enzyme inhibitors. NRPS has a distinct modular structure in which each module is responsible for the recognition, activation, and in some cases, modification of a single amino acid residue of the final peptide product. The modules can be subdivided into domains that catalyze specific biochemical reactions. This family includes the myxovirescin (TA) antibiotic biosynthetic gene in Myxococcus xanthus; TA production plays a role in predation. It also includes the salinosporamide A polyketide synthase which is involved in the biosynthesis of salinosporamide A, a marine microbial metabolite whose chlorine atom is crucial for potent proteasome inhibition and anticancer activity. |
| cd05930 | A_NRPS | 1.11e-159 | 582 | 998 | 3 | 395 | The adenylation domain of nonribosomal peptide synthetases (NRPS). The adenylation (A) domain of NRPS recognizes a specific amino acid or hydroxy acid and activates it as an (amino) acyl adenylate by hydrolysis of ATP. The activated acyl moiety then forms a thioester bond to the enzyme-bound cofactor phosphopantetheine of a peptidyl carrier protein domain. NRPSs are large multifunctional enzymes which synthesize many therapeutically useful peptides in bacteria and fungi via a template-directed, nucleic acid independent nonribosomal mechanism. These natural products include antibiotics, immunosuppressants, plant and animal toxins, and enzyme inhibitors. NRPS has a distinct modular structure in which each module is responsible for the recognition, activation, and in some cases, modification of a single amino acid residue of the final peptide product. The modules can be subdivided into domains that catalyze specific biochemical reactions. |
| smart00825 | PKS_KS | 2.07e-157 | 1552 | 1976 | 1 | 298 | Beta-ketoacyl synthase. The structure of beta-ketoacyl synthase is similar to that of the thiolase family and also chalcone synthase. The active site of beta-ketoacyl synthase is located between the N and C-terminal domains. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFY93865.1 | 4.41e-193 | 540 | 2443 | 305 | 1867 |
| BAY90071.1 | 2.83e-178 | 531 | 2445 | 543 | 2102 |
| BAY30132.1 | 9.86e-155 | 1410 | 2439 | 1043 | 2098 |
| BAZ00088.1 | 3.80e-154 | 1410 | 2439 | 1043 | 2096 |
| BAZ75991.1 | 3.80e-154 | 1410 | 2439 | 1043 | 2096 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4MZ0_A | 8.58e-251 | 1535 | 2438 | 24 | 937 | ChainA, CurL [Moorena producens 3L],4MZ0_B Chain B, CurL [Moorena producens 3L] |
| 2HG4_A | 7.61e-205 | 1548 | 2442 | 37 | 903 | Structureof the ketosynthase-acyltransferase didomain of module 5 from DEBS. [Saccharopolyspora erythraea],2HG4_B Structure of the ketosynthase-acyltransferase didomain of module 5 from DEBS. [Saccharopolyspora erythraea],2HG4_C Structure of the ketosynthase-acyltransferase didomain of module 5 from DEBS. [Saccharopolyspora erythraea],2HG4_D Structure of the ketosynthase-acyltransferase didomain of module 5 from DEBS. [Saccharopolyspora erythraea],2HG4_E Structure of the ketosynthase-acyltransferase didomain of module 5 from DEBS. [Saccharopolyspora erythraea],2HG4_F Structure of the ketosynthase-acyltransferase didomain of module 5 from DEBS. [Saccharopolyspora erythraea] |
| 7VEE_A | 5.43e-204 | 1548 | 2445 | 19 | 920 | ChainA, Polyketide synthase [Streptomyces graminofaciens],7VEF_A Chain A, Polyketide synthase [Streptomyces graminofaciens] |
| 7M7J_A | 1.35e-203 | 1542 | 2441 | 25 | 914 | ChainA, EryAI [Saccharopolyspora erythraea],7M7J_B Chain B, EryAI [Saccharopolyspora erythraea] |
| 7M7I_A | 1.35e-203 | 1542 | 2441 | 25 | 914 | ChainA, EryAI [Saccharopolyspora erythraea],7M7I_B Chain B, EryAI [Saccharopolyspora erythraea] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D4AU31 | 0.0 | 542 | 4023 | 9 | 2558 | PKS-NRPS hybrid synthetase swnK OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=swnK PE=3 SV=1 |
| Q03132 | 0.0 | 1454 | 3621 | 1388 | 3488 | 6-deoxyerythronolide-B synthase EryA2, modules 3 and 4 OS=Saccharopolyspora erythraea OX=1836 GN=eryA PE=1 SV=3 |
| B2HIL7 | 0.0 | 1534 | 3634 | 20 | 2088 | Phenolphthiocerol synthesis polyketide synthase type I Pks15/1 OS=Mycobacterium marinum (strain ATCC BAA-535 / M) OX=216594 GN=pks15/1 PE=1 SV=1 |
| Q7TXK8 | 0.0 | 1534 | 3622 | 25 | 2086 | Phenolphthiocerol synthesis polyketide synthase type I Pks15/1 OS=Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97) OX=233413 GN=pks15/1 PE=1 SV=1 |
| E9F8M3 | 0.0 | 540 | 4026 | 1 | 2482 | PKS-NRPS hybrid synthetase swnK OS=Metarhizium robertsii (strain ARSEF 23 / ATCC MYA-3075) OX=655844 GN=swnK PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
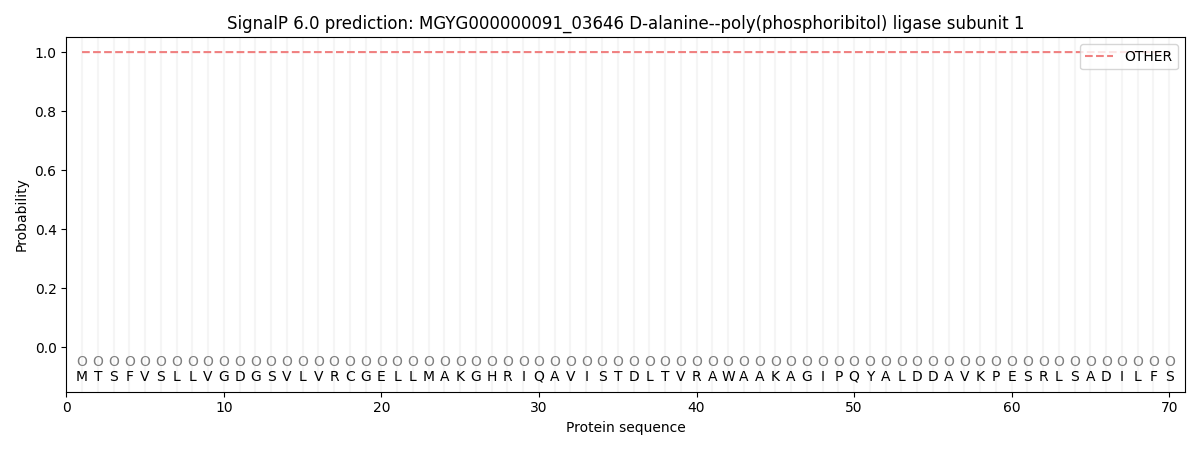
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000042 | 0.000003 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
