You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000091_06058
You are here: Home > Sequence: MGYG000000091_06058
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Achromobacter xylosoxidans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Burkholderiales; Burkholderiaceae; Achromobacter; Achromobacter xylosoxidans | |||||||||||
| CAZyme ID | MGYG000000091_06058 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | Peptidoglycan hydrolase FlgJ | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 98171; End: 99154 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK12713 | flgJ | 2.83e-169 | 1 | 320 | 1 | 336 | flagellar rod assembly protein/muramidase FlgJ; Provisional |
| TIGR02541 | flagell_FlgJ | 3.66e-128 | 16 | 311 | 1 | 294 | flagellar rod assembly protein/muramidase FlgJ. The N-terminal region of this protein acts directly in flagellar rod assembly. The C-terminal region is a flagellum-specific muramidase (peptidoglycan hydrolase) required for formation of the outer membrane L ring. |
| PRK05684 | flgJ | 4.31e-114 | 18 | 317 | 12 | 304 | flagellar assembly peptidoglycan hydrolase FlgJ. |
| PRK12712 | flgJ | 1.45e-87 | 35 | 310 | 34 | 342 | flagellar rod assembly protein/muramidase FlgJ; Provisional |
| PRK12709 | flgJ | 1.54e-77 | 35 | 310 | 30 | 319 | flagellar rod assembly protein/muramidase FlgJ; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CCH08218.1 | 1.50e-229 | 1 | 327 | 1 | 327 |
| CKH30351.1 | 1.50e-229 | 1 | 327 | 1 | 327 |
| QKQ57085.1 | 1.50e-229 | 1 | 327 | 1 | 327 |
| QEQ23009.1 | 1.50e-229 | 1 | 327 | 1 | 327 |
| SQG71987.1 | 1.50e-229 | 1 | 327 | 1 | 327 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5DN4_A | 7.83e-52 | 167 | 318 | 5 | 156 | Structureof the glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 5DN5_A | 1.09e-50 | 167 | 311 | 5 | 149 | Structureof a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_B Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_C Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 3VWO_A | 9.34e-43 | 168 | 308 | 4 | 144 | Crystalstructure of peptidoglycan hydrolase mutant from Sphingomonas sp. A1 [Sphingomonas sp. A1] |
| 2ZYC_A | 1.22e-42 | 168 | 308 | 5 | 145 | ChainA, Peptidoglycan hydrolase FlgJ [Sphingomonas sp. A1] |
| 3K3T_A | 9.57e-42 | 168 | 308 | 5 | 145 | E185Amutant of peptidoglycan hydrolase from Sphingomonas sp. A1 [Sphingomonas sp. A1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P15931 | 1.39e-71 | 16 | 318 | 11 | 305 | Peptidoglycan hydrolase FlgJ OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=flgJ PE=1 SV=1 |
| P58231 | 3.59e-71 | 16 | 323 | 11 | 307 | Peptidoglycan hydrolase FlgJ OS=Escherichia coli O157:H7 OX=83334 GN=flgJ PE=3 SV=1 |
| P75942 | 2.86e-70 | 16 | 323 | 11 | 307 | Peptidoglycan hydrolase FlgJ OS=Escherichia coli (strain K12) OX=83333 GN=flgJ PE=3 SV=1 |
| Q9X9J3 | 1.43e-45 | 17 | 310 | 11 | 305 | Peptidoglycan hydrolase FlgJ OS=Vibrio parahaemolyticus serotype O3:K6 (strain RIMD 2210633) OX=223926 GN=flgJ PE=3 SV=1 |
| Q9KQ15 | 3.13e-43 | 5 | 310 | 3 | 306 | Peptidoglycan hydrolase FlgJ OS=Vibrio cholerae serotype O1 (strain ATCC 39315 / El Tor Inaba N16961) OX=243277 GN=flgJ PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
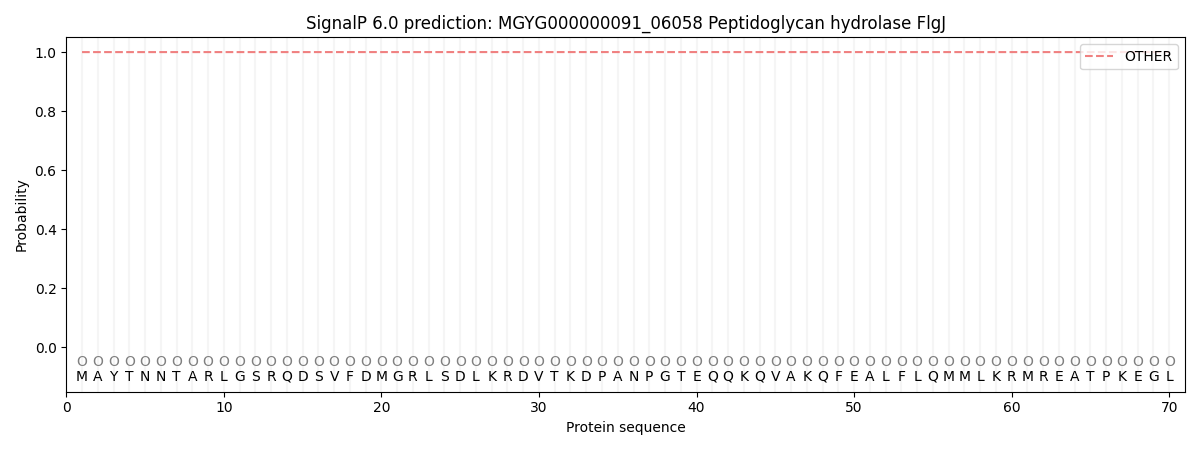
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000053 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
