You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000097_01829
You are here: Home > Sequence: MGYG000000097_01829
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
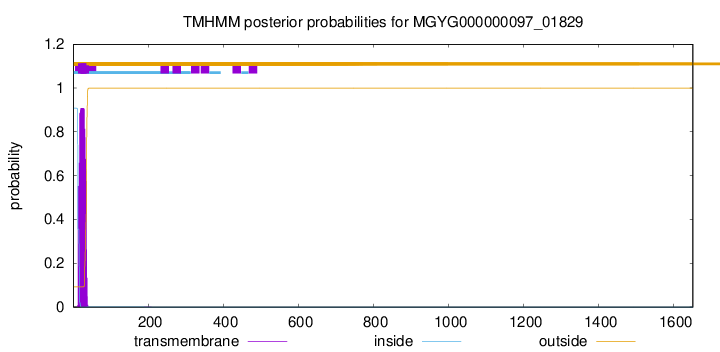
TMHMM annotations
Basic Information help
| Species | Ligilactobacillus animalis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Ligilactobacillus; Ligilactobacillus animalis | |||||||||||
| CAZyme ID | MGYG000000097_01829 | |||||||||||
| CAZy Family | GH70 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4920; End: 9875 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH70 | 728 | 1526 | 0 | 0.9975093399750934 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02324 | Glyco_hydro_70 | 0.0 | 728 | 1527 | 1 | 803 | Glycosyl hydrolase family 70. Members of this family belong to glycosyl hydrolase family 70 Glucosyltransferases or sucrose 6-glycosyl transferases (GTF-S) catalyze the transfer of D-glucopyramnosyl units from sucrose onto acceptor molecules, EC:2.4.1.5. This family roughly corresponds to the N-terminal catalytic domain of the enzyme. Members of this family also contain the Putative cell wall binding domain pfam01473, which corresponds with the C-terminal glucan-binding domain. |
| 227588 | COG5263 | 1.93e-22 | 325 | 521 | 139 | 309 | COG5263 Glucan-binding domain (YG repeat) [Carbohydrate transport and metabolism]. |
| NF033930 | pneumo_PspA | 3.11e-21 | 346 | 546 | 452 | 619 | pneumococcal surface protein A. The pneumococcal surface protein proteins, found in Streptococcus pneumoniae, are repetitive, with patterns of localized high sequence identity across pairs of proteins given different specific names that recombination may be presumed. This protein, PspA, has an N-terminal region that lacks a cross-wall-targeting YSIRK type extended signal peptide, in contrast to the closely related choline-binding protein CbpA which has a similar C-terminus but a YSIRK-containing region at the N-terminus. |
| NF033930 | pneumo_PspA | 4.30e-20 | 381 | 552 | 444 | 605 | pneumococcal surface protein A. The pneumococcal surface protein proteins, found in Streptococcus pneumoniae, are repetitive, with patterns of localized high sequence identity across pairs of proteins given different specific names that recombination may be presumed. This protein, PspA, has an N-terminal region that lacks a cross-wall-targeting YSIRK type extended signal peptide, in contrast to the closely related choline-binding protein CbpA which has a similar C-terminus but a YSIRK-containing region at the N-terminus. |
| NF033930 | pneumo_PspA | 1.55e-18 | 401 | 679 | 443 | 660 | pneumococcal surface protein A. The pneumococcal surface protein proteins, found in Streptococcus pneumoniae, are repetitive, with patterns of localized high sequence identity across pairs of proteins given different specific names that recombination may be presumed. This protein, PspA, has an N-terminal region that lacks a cross-wall-targeting YSIRK type extended signal peptide, in contrast to the closely related choline-binding protein CbpA which has a similar C-terminus but a YSIRK-containing region at the N-terminus. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QHQ69356.1 | 0.0 | 1 | 1651 | 1 | 1651 |
| CCK33644.1 | 0.0 | 1 | 1651 | 1 | 1585 |
| QHQ69357.1 | 0.0 | 2 | 1631 | 4 | 1572 |
| AUJ33173.1 | 0.0 | 642 | 1537 | 292 | 1195 |
| AYP74780.1 | 0.0 | 637 | 1594 | 752 | 1708 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3KLL_A | 0.0 | 637 | 1594 | 18 | 974 | Crystalstructure of Lactobacillus reuteri N-terminally truncated glucansucrase GTF180-maltose complex [Limosilactobacillus reuteri] |
| 4AMC_A | 0.0 | 637 | 1597 | 18 | 985 | Crystalstructure of Lactobacillus reuteri 121 N-terminally truncated glucansucrase GTFA [Limosilactobacillus reuteri] |
| 3HZ3_A | 0.0 | 637 | 1594 | 18 | 974 | Lactobacillusreuteri N-terminally truncated glucansucrase GTF180(D1025N)-sucrose complex [Limosilactobacillus reuteri] |
| 3KLK_A | 0.0 | 637 | 1594 | 18 | 974 | Crystalstructure of Lactobacillus reuteri N-terminally truncated glucansucrase GTF180 in triclinic apo- form [Limosilactobacillus reuteri],4AYG_A Lactobacillus reuteri N-terminally truncated glucansucrase GTF180 in orthorhombic apo-form [Limosilactobacillus reuteri],4AYG_B Lactobacillus reuteri N-terminally truncated glucansucrase GTF180 in orthorhombic apo-form [Limosilactobacillus reuteri] |
| 6SYQ_B | 5.37e-293 | 646 | 1600 | 133 | 1153 | ChainB, Alternansucrase [Leuconostoc mesenteroides] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P49331 | 4.93e-276 | 637 | 1603 | 191 | 1150 | Glucosyltransferase-S OS=Streptococcus mutans serotype c (strain ATCC 700610 / UA159) OX=210007 GN=gtfD PE=3 SV=3 |
| P08987 | 2.17e-265 | 636 | 1609 | 178 | 1148 | Glucosyltransferase-I OS=Streptococcus mutans serotype c (strain ATCC 700610 / UA159) OX=210007 GN=gtfB PE=3 SV=3 |
| P13470 | 1.73e-260 | 668 | 1609 | 232 | 1177 | Glucosyltransferase-SI OS=Streptococcus mutans serotype c (strain ATCC 700610 / UA159) OX=210007 GN=gtfC PE=1 SV=2 |
| P11001 | 8.41e-258 | 635 | 1628 | 175 | 1169 | Glucosyltransferase-I OS=Streptococcus downei OX=1317 GN=gtfI PE=3 SV=1 |
| P27470 | 1.03e-257 | 635 | 1609 | 169 | 1144 | Glucosyltransferase-I OS=Streptococcus downei OX=1317 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
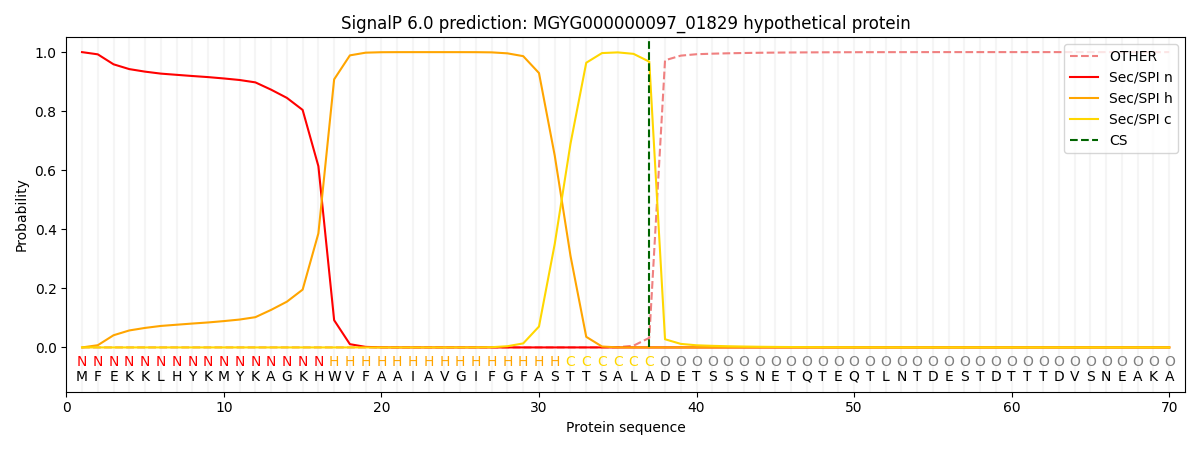
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000621 | 0.998534 | 0.000287 | 0.000188 | 0.000170 | 0.000154 |