You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000102_03381
You are here: Home > Sequence: MGYG000000102_03381
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Terrisporobacter sp902363255 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Peptostreptococcales; Peptostreptococcaceae; Terrisporobacter; Terrisporobacter sp902363255 | |||||||||||
| CAZyme ID | MGYG000000102_03381 | |||||||||||
| CAZy Family | GH0 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 41254; End: 42525 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01471 | PG_binding_1 | 2.90e-14 | 349 | 406 | 5 | 57 | Putative peptidoglycan binding domain. This domain is composed of three alpha helices. This domain is found at the N or C-terminus of a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. This family is found N-terminal to the catalytic domain of matrixins. The domain is found to bind peptidoglycan experimentally. |
| COG3409 | PGRP | 7.57e-09 | 319 | 408 | 100 | 185 | Peptidoglycan-binding (PGRP) domain of peptidoglycan hydrolases [Cell wall/membrane/envelope biogenesis]. |
| COG3409 | PGRP | 1.13e-06 | 349 | 406 | 48 | 102 | Peptidoglycan-binding (PGRP) domain of peptidoglycan hydrolases [Cell wall/membrane/envelope biogenesis]. |
| COG2385 | SpoIID | 0.005 | 293 | 316 | 362 | 385 | Peptidoglycan hydrolase (amidase) enhancer domain [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AXU85473.1 | 2.23e-238 | 1 | 423 | 1 | 423 |
| QOX88424.1 | 2.23e-238 | 1 | 423 | 1 | 423 |
| AXU70569.1 | 2.23e-238 | 1 | 423 | 1 | 423 |
| CDS83713.1 | 2.23e-238 | 1 | 423 | 1 | 423 |
| CAJ67384.1 | 2.23e-238 | 1 | 423 | 1 | 423 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
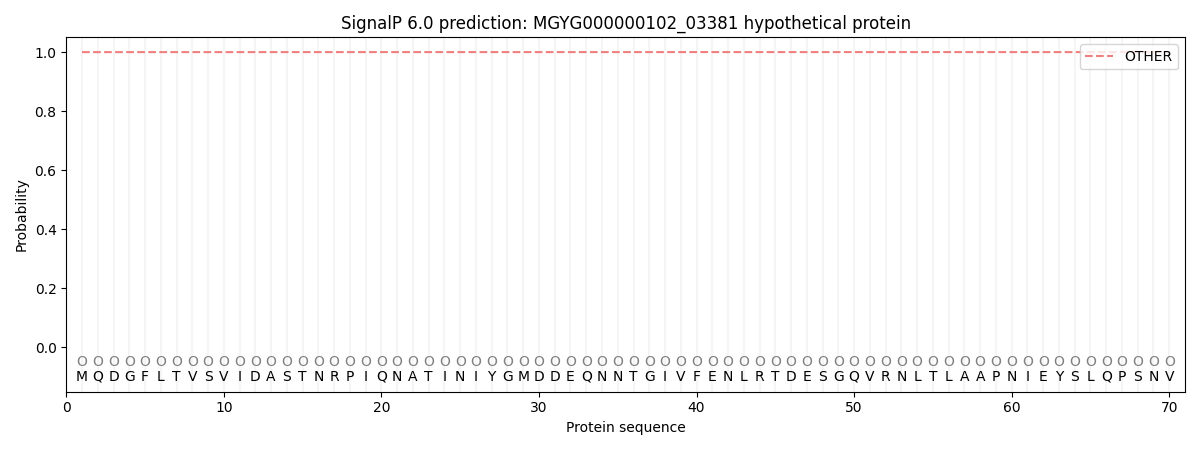
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000003 | 0.000021 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
