You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000113_01460
You are here: Home > Sequence: MGYG000000113_01460
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
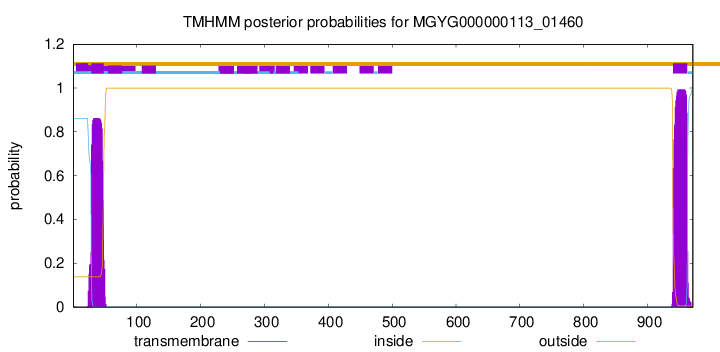
TMHMM annotations
Basic Information help
| Species | Streptococcus salivarius | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus salivarius | |||||||||||
| CAZyme ID | MGYG000000113_01460 | |||||||||||
| CAZy Family | GH68 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 121772; End: 124687 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH68 | 232 | 684 | 1.1e-138 | 0.988009592326139 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02435 | Glyco_hydro_68 | 1.17e-116 | 231 | 675 | 2 | 410 | Levansucrase/Invertase. This Pfam family consists of the glycosyl hydrolase 68 family, including several bacterial levansucrase enzymes, and invertase from zymomonas. |
| cd08997 | GH68 | 1.62e-99 | 285 | 672 | 1 | 352 | Glycosyl hydrolase family 68, includes levansucrase, beta-fructofuranosidase and inulosucrase. Glycosyl hydrolase family 68 (GH68) consists of frucosyltransferases (FTFs) that include levansucrase (EC 2.4.1.10), beta-fructofuranosidase (EC 3.2.1.26) and inulosucrase (EC 2.4.1.9), all of which use sucrose as their preferential donor substrate. Levansucrase, also known as beta-D-fructofuranosyl transferase, catalyzes the transfer of the sucrose fructosyl moiety to a growing levan chain. Similarly, inulosucrase catalyzes long inulin-type of fructans, and beta-fructofuranosidases create fructooligosaccharides (FOS). However, in the absence of high fructan/sucrose ratio, some GH68 enzymes can also use fructan as donor substrate. GH68 retaining enzymes (i.e. they retain the configuration at anomeric carbon atom of the substrate) catalyze hydrolysis in two steps involving a covalent glycosyl enzyme intermediate: an aspartate located close to the N-terminus acts as the catalytic nucleophile and a glutamate acts as the general acid/base; a conserved aspartate residue in the Arg-Asp-Pro (RDP) motif stabilizes the transition state. A common structural feature of all these enzymes is a 5-bladed beta-propeller domain, similar to GH43, that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. Biotechnological applications of these enzymes include use of inulin in inexpensive production of rich fructose syrups as well as use of FOS as health-promoting pre-biotics. |
| 350093 | cd08979 | 7.18e-23 | 286 | 664 | 1 | 291 | GH_J Glycosyl hydrolase families 32 and 68, which form the clan GH-J. This glycosyl hydrolase family clan J (according to carbohydrate-active enzymes database (CAZY)) includes family 32 (GH32) and 68 (GH68). GH32 enzymes include invertase (EC 3.2.1.26) and other other fructofuranosidases such as inulinase (EC 3.2.1.7), exo-inulinase (EC 3.2.1.80), levanase (EC 3.2.1.65), and transfructosidases such sucrose:sucrose 1-fructosyltransferase (EC 2.4.1.99), fructan:fructan 1-fructosyltransferase (EC 2.4.1.100), sucrose:fructan 6-fructosyltransferase (EC 2.4.1.10), fructan:fructan 6G-fructosyltransferase (EC 2.4.1.243) and levan fructosyltransferases (EC 2.4.1.-). The GH68 family consists of frucosyltransferases (FTFs) that include levansucrase (EC 2.4.1.10, also known as beta-D-fructofuranosyl transferase), beta-fructofuranosidase (EC 3.2.1.26) and inulosucrase (EC 2.4.1.9). GH32 and GH68 family enzymes are retaining enzymes (i.e. they retain the configuration at anomeric carbon atom of the substrate) and catalyze hydrolysis in two steps involving a covalent glycosyl enzyme intermediate: an aspartate located close to the N-terminus acts as the catalytic nucleophile and a glutamate acts as the general acid/base; a conserved aspartate residue in the Arg-Asp-Pro (RDP) motif stabilizes the transition state. A common structural feature of all these enzymes is a 5-bladed beta-propeller domain, similar to GH43, that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| PRK10811 | rne | 1.49e-08 | 47 | 203 | 861 | 1021 | ribonuclease E; Reviewed |
| PRK10811 | rne | 7.74e-08 | 31 | 189 | 863 | 1022 | ribonuclease E; Reviewed |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ARC33740.1 | 0.0 | 1 | 971 | 1 | 971 |
| QGU79950.1 | 0.0 | 1 | 971 | 1 | 971 |
| ALR79294.1 | 0.0 | 1 | 971 | 1 | 971 |
| CCB94680.1 | 0.0 | 1 | 971 | 1 | 971 |
| AMB82148.1 | 0.0 | 1 | 971 | 1 | 971 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2YFR_A | 3.18e-207 | 185 | 726 | 31 | 566 | Crystalstructure of inulosucrase from Lactobacillus johnsonii NCC533 [Lactobacillus johnsonii],2YFT_A Crystal structure of inulosucrase from Lactobacillus johnsonii NCC533 in complex with 1-kestose [Lactobacillus johnsonii] |
| 2YFS_A | 1.79e-206 | 185 | 726 | 31 | 566 | Crystalstructure of inulosucrase from Lactobacillus johnsonii NCC533 in complex with sucrose [Lactobacillus johnsonii] |
| 3OM6_A | 1.56e-87 | 249 | 682 | 30 | 449 | ChainA, Levansucrase [Priestia megaterium],3OM6_B Chain B, Levansucrase [Priestia megaterium],3OM6_C Chain C, Levansucrase [Priestia megaterium],3OM6_D Chain D, Levansucrase [Priestia megaterium] |
| 3OM4_A | 3.01e-87 | 249 | 682 | 30 | 449 | ChainA, Levansucrase [Priestia megaterium],3OM4_B Chain B, Levansucrase [Priestia megaterium],3OM4_C Chain C, Levansucrase [Priestia megaterium],3OM4_D Chain D, Levansucrase [Priestia megaterium] |
| 3OM7_A | 4.19e-87 | 249 | 682 | 30 | 449 | ChainA, Levansucrase [Priestia megaterium],3OM7_B Chain B, Levansucrase [Priestia megaterium],3OM7_C Chain C, Levansucrase [Priestia megaterium],3OM7_D Chain D, Levansucrase [Priestia megaterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q55242 | 0.0 | 1 | 971 | 1 | 969 | Levansucrase OS=Streptococcus salivarius OX=1304 GN=ftf PE=3 SV=1 |
| P11701 | 3.85e-204 | 209 | 725 | 170 | 678 | Levansucrase OS=Streptococcus mutans serotype c (strain ATCC 700610 / UA159) OX=210007 GN=ftf PE=3 SV=2 |
| Q74K42 | 4.59e-203 | 185 | 724 | 174 | 707 | Inulosucrase OS=Lactobacillus johnsonii (strain CNCM I-12250 / La1 / NCC 533) OX=257314 GN=inuJ PE=1 SV=1 |
| D3WYV9 | 5.68e-200 | 186 | 724 | 169 | 701 | Inulosucrase OS=Lactobacillus gasseri OX=1596 GN=inuGB PE=1 SV=1 |
| D3WYW0 | 2.00e-199 | 188 | 724 | 156 | 688 | Levansucrase OS=Lactobacillus gasseri OX=1596 GN=levG PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.045793 | 0.947649 | 0.004853 | 0.001063 | 0.000391 | 0.000248 |