You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000123_02537
You are here: Home > Sequence: MGYG000000123_02537
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Blautia sp001304935 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Blautia; Blautia sp001304935 | |||||||||||
| CAZyme ID | MGYG000000123_02537 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | Tyrocidine synthase 3 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 171737; End: 179209 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK05691 | PRK05691 | 0.0 | 42 | 2091 | 674 | 2769 | peptide synthase; Validated |
| PRK12316 | PRK12316 | 0.0 | 115 | 2090 | 1627 | 3615 | peptide synthase; Provisional |
| PRK12467 | PRK12467 | 0.0 | 28 | 2092 | 26 | 2158 | peptide synthase; Provisional |
| PRK05691 | PRK05691 | 1.45e-172 | 489 | 2070 | 2 | 1677 | peptide synthase; Validated |
| PRK12316 | PRK12316 | 1.95e-172 | 1046 | 2091 | 22 | 1078 | peptide synthase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QND46664.1 | 5.32e-198 | 145 | 2075 | 665 | 2639 |
| ACX49739.1 | 6.72e-134 | 44 | 1819 | 10 | 1880 |
| BAY90071.1 | 6.05e-115 | 980 | 2066 | 2131 | 3251 |
| BAY30132.1 | 8.08e-113 | 980 | 2066 | 2142 | 3262 |
| BAZ00088.1 | 1.06e-112 | 980 | 2066 | 2140 | 3260 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6MFZ_A | 9.78e-203 | 505 | 2094 | 213 | 1793 | Crystalstructure of dimodular LgrA in a condensation state [Brevibacillus parabrevis],6MFZ_B Crystal structure of dimodular LgrA in a condensation state [Brevibacillus parabrevis] |
| 6MFY_A | 2.05e-198 | 505 | 1996 | 213 | 1712 | Crystalstructure of a 5-domain construct of LgrA in the substrate donation state [Brevibacillus parabrevis],6MG0_A Crystal structure of a 5-domain construct of LgrA in the thiolation state [Brevibacillus parabrevis],6MG0_B Crystal structure of a 5-domain construct of LgrA in the thiolation state [Brevibacillus parabrevis] |
| 5U89_A | 6.38e-125 | 476 | 1501 | 4 | 1071 | Crystalstructure of a cross-module fragment from the dimodular NRPS DhbF [Geobacillus sp. Y4.1MC1] |
| 6MFX_A | 1.85e-117 | 505 | 1498 | 213 | 1207 | Crystalstructure of a 4-domain construct of a mutant of LgrA in the substrate donation state [Brevibacillus parabrevis] |
| 6MFW_A | 2.49e-117 | 505 | 1498 | 213 | 1207 | Crystalstructure of a 4-domain construct of LgrA in the substrate donation state [Brevibacillus parabrevis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39845 | 5.98e-284 | 46 | 2102 | 8 | 2085 | Plipastatin synthase subunit A OS=Bacillus subtilis (strain 168) OX=224308 GN=ppsA PE=1 SV=2 |
| P27206 | 5.60e-279 | 43 | 2142 | 5 | 2125 | Surfactin synthase subunit 1 OS=Bacillus subtilis (strain 168) OX=224308 GN=srfAA PE=1 SV=4 |
| O30409 | 1.88e-278 | 35 | 2070 | 2 | 2045 | Tyrocidine synthase 3 OS=Brevibacillus parabrevis OX=54914 GN=tycC PE=1 SV=1 |
| P0C064 | 9.97e-273 | 41 | 2097 | 2094 | 4160 | Gramicidin S synthase 2 OS=Brevibacillus brevis OX=1393 GN=grsB PE=1 SV=2 |
| P94459 | 3.02e-265 | 34 | 2100 | 1047 | 3107 | Plipastatin synthase subunit D OS=Bacillus subtilis (strain 168) OX=224308 GN=ppsD PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
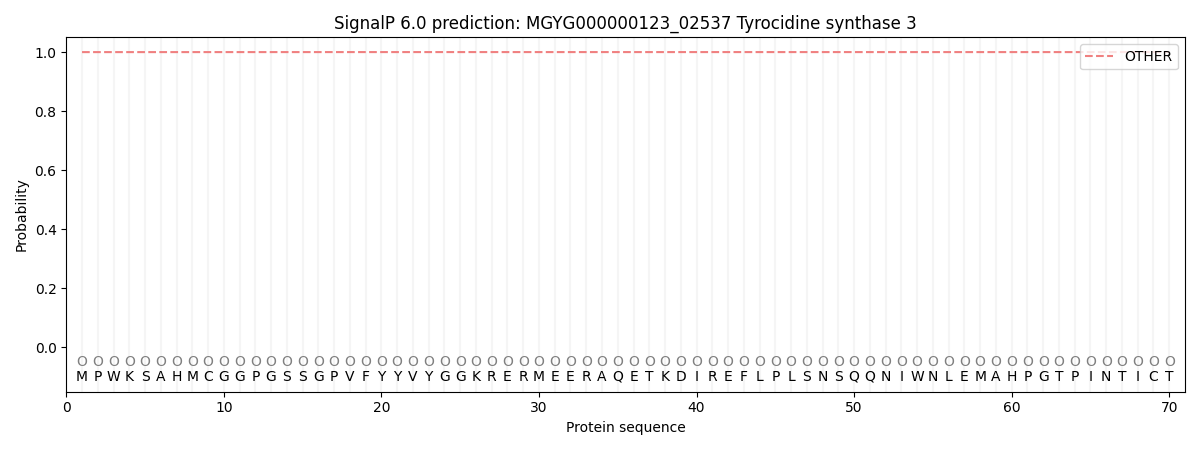
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000046 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
