You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000125_00590
You are here: Home > Sequence: MGYG000000125_00590
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Clostridium_B tyrobutyricum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium_B; Clostridium_B tyrobutyricum | |||||||||||
| CAZyme ID | MGYG000000125_00590 | |||||||||||
| CAZy Family | GT83 | |||||||||||
| CAZyme Description | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 91719; End: 93788 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT83 | 12 | 229 | 1.9e-33 | 0.39814814814814814 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1807 | ArnT | 1.84e-70 | 6 | 651 | 2 | 531 | 4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferase of PMT family [Cell wall/membrane/envelope biogenesis]. |
| pfam13231 | PMT_2 | 4.60e-41 | 67 | 225 | 1 | 160 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This family contains members that are not captured by pfam02366. |
| COG1928 | PMT1 | 7.89e-04 | 1 | 183 | 14 | 211 | Dolichyl-phosphate-mannose--protein O-mannosyl transferase [Posttranslational modification, protein turnover, chaperones]. |
| pfam02516 | STT3 | 0.001 | 58 | 224 | 57 | 235 | Oligosaccharyl transferase STT3 subunit. This family consists of the oligosaccharyl transferase STT3 subunit and related proteins. The STT3 subunit is part of the oligosaccharyl transferase (OTase) complex of proteins and is required for its activity. In eukaryotes, OTase transfers a lipid-linked core-oligosaccharide to selected asparagine residues in the ER. In the archaea STT3 occurs alone, rather than in an OTase complex, and is required for N-glycosylation of asparagines. |
| COG4745 | COG4745 | 0.002 | 81 | 231 | 78 | 231 | Predicted membrane-bound mannosyltransferase [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUB37647.1 | 2.43e-77 | 17 | 646 | 210 | 780 |
| QHU89858.1 | 6.68e-76 | 32 | 646 | 293 | 889 |
| QHU90800.1 | 8.03e-74 | 32 | 646 | 293 | 886 |
| QLF52252.1 | 2.75e-71 | 32 | 646 | 293 | 891 |
| QJU11362.1 | 2.75e-71 | 32 | 646 | 293 | 891 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5EZM_A | 3.50e-06 | 67 | 230 | 87 | 255 | CrystalStructure of ArnT from Cupriavidus metallidurans in the apo state [Cupriavidus metallidurans CH34],5F15_A Crystal Structure of ArnT from Cupriavidus metallidurans bound to Undecaprenyl phosphate [Cupriavidus metallidurans CH34] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34575 | 3.77e-120 | 12 | 663 | 10 | 679 | Putative mannosyltransferase YkcB OS=Bacillus subtilis (strain 168) OX=224308 GN=ykcB PE=3 SV=2 |
| P37483 | 3.19e-116 | 8 | 657 | 3 | 662 | Putative mannosyltransferase YycA OS=Bacillus subtilis (strain 168) OX=224308 GN=yycA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
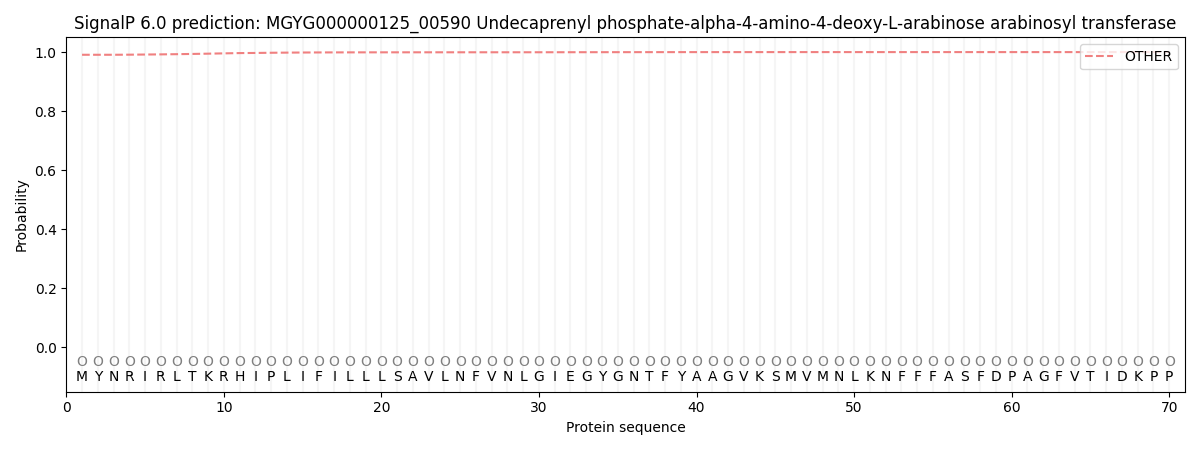
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.991005 | 0.000909 | 0.000195 | 0.000006 | 0.000004 | 0.007907 |
TMHMM Annotations download full data without filtering help
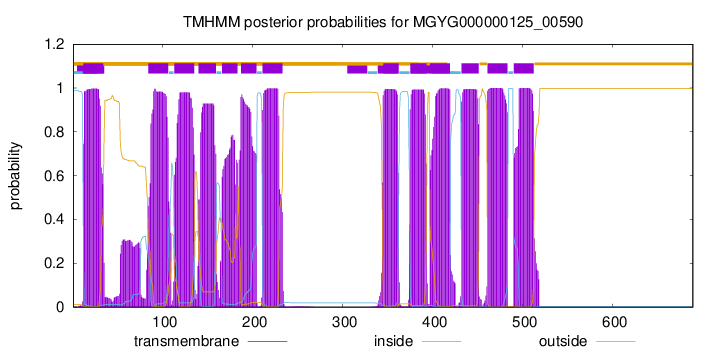
| start | end |
|---|---|
| 12 | 34 |
| 84 | 106 |
| 113 | 135 |
| 140 | 159 |
| 166 | 183 |
| 187 | 204 |
| 211 | 233 |
| 345 | 362 |
| 375 | 393 |
| 397 | 419 |
| 432 | 451 |
| 461 | 483 |
| 490 | 512 |
