You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000135_04484
You are here: Home > Sequence: MGYG000000135_04484
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Murimonas intestini | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Murimonas; Murimonas intestini | |||||||||||
| CAZyme ID | MGYG000000135_04484 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-N-acetylglucosaminidase/beta-glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 21484; End: 23181 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 78 | 312 | 3.5e-51 | 0.9768518518518519 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 5.19e-57 | 26 | 417 | 1 | 361 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 8.99e-56 | 29 | 348 | 3 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK05337 | PRK05337 | 7.29e-26 | 117 | 316 | 96 | 282 | beta-hexosaminidase; Provisional |
| PRK15098 | PRK15098 | 2.85e-16 | 6 | 402 | 26 | 407 | beta-glucosidase BglX. |
| cd13643 | PBP2_BCP_2 | 0.008 | 205 | 253 | 47 | 102 | Substrate-binding domain of osmoregulatory ABC-type glycine betaine/choline/L-proline transport system-like; the type 2 periplasmic-binding protein fold. This subfamily is part of a high affinity multicomponent binding-protein-dependent transport system specific to certain quaternary ammonium compounds for osmoregulation. The periplasmic substrate-binding domain, which is often fused to the permease component of the ATP-binding cassette transporter complex, is involved in uptake of osmoprotectants (also termed compatible solutes) such as glycine betaine, proline betaine, choline, and carnitine. Many microorganisms accumulate these compatible solutes in response to high osmolarity to offset the loss of cell water. This domain belongs to the type 2 periplasmic binding fold protein superfamily (PBP2). The PBP2 proteins are typically comprised of two globular subdomains connected by a flexible hinge and bind their ligand in the cleft between these domains in a manner resembling a Venus flytrap. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AQM60421.1 | 1.32e-304 | 1 | 565 | 1 | 565 |
| QMW79114.1 | 1.61e-273 | 1 | 565 | 1 | 569 |
| QIB54599.1 | 1.61e-273 | 1 | 565 | 1 | 569 |
| QBE97674.1 | 1.32e-272 | 1 | 565 | 1 | 569 |
| QTQ13616.1 | 1.58e-266 | 1 | 565 | 1 | 569 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5VQD_A | 1.83e-131 | 4 | 565 | 6 | 567 | Beta-glucosidephosphorylase BglX [unidentified],5VQE_A Beta-glucoside phosphorylase BglX bound to 2FGlc [unidentified] |
| 3BMX_A | 2.80e-42 | 26 | 413 | 42 | 458 | Beta-N-hexosaminidase(YbbD) from Bacillus subtilis [Bacillus subtilis],3BMX_B Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3NVD_A Structure of YBBD in complex with pugnac [Bacillus subtilis],3NVD_B Structure of YBBD in complex with pugnac [Bacillus subtilis] |
| 3LK6_A | 1.01e-41 | 26 | 413 | 16 | 432 | ChainA, Lipoprotein ybbD [Bacillus subtilis],3LK6_B Chain B, Lipoprotein ybbD [Bacillus subtilis],3LK6_C Chain C, Lipoprotein ybbD [Bacillus subtilis],3LK6_D Chain D, Lipoprotein ybbD [Bacillus subtilis] |
| 4GYJ_A | 1.39e-41 | 26 | 413 | 46 | 462 | Crystalstructure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYJ_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYK_A Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168],4GYK_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168] |
| 6K5J_A | 1.53e-37 | 29 | 550 | 14 | 525 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q7WUL3 | 6.73e-138 | 1 | 565 | 1 | 564 | Beta-N-acetylglucosaminidase/beta-glucosidase OS=Cellulomonas fimi OX=1708 GN=nag3 PE=1 SV=1 |
| P40406 | 1.53e-41 | 26 | 413 | 42 | 458 | Beta-hexosaminidase OS=Bacillus subtilis (strain 168) OX=224308 GN=nagZ PE=1 SV=1 |
| Q7NWB7 | 2.12e-24 | 114 | 319 | 94 | 290 | Beta-hexosaminidase OS=Chromobacterium violaceum (strain ATCC 12472 / DSM 30191 / JCM 1249 / NBRC 12614 / NCIMB 9131 / NCTC 9757) OX=243365 GN=nagZ PE=3 SV=1 |
| A9M1Z4 | 2.53e-23 | 113 | 321 | 100 | 302 | Beta-hexosaminidase OS=Neisseria meningitidis serogroup C (strain 053442) OX=374833 GN=nagZ PE=3 SV=1 |
| Q1H075 | 2.87e-23 | 116 | 319 | 97 | 288 | Beta-hexosaminidase OS=Methylobacillus flagellatus (strain KT / ATCC 51484 / DSM 6875) OX=265072 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
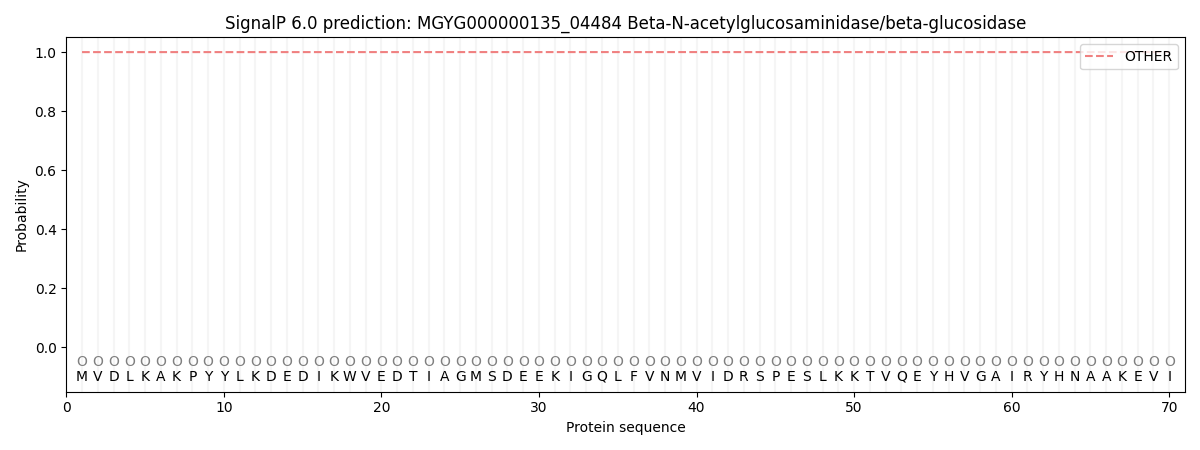
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000028 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
