You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000139_01646
You are here: Home > Sequence: MGYG000000139_01646
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
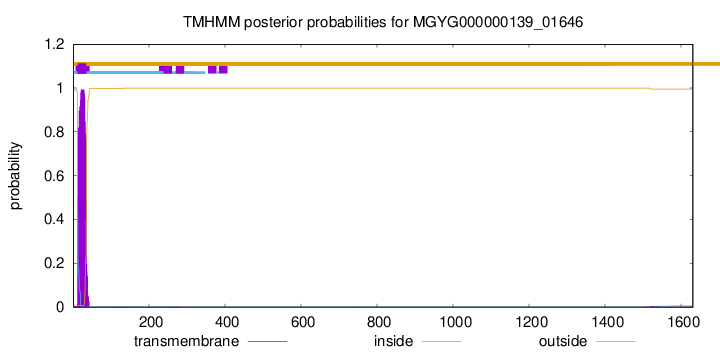
TMHMM annotations
Basic Information help
| Species | Eubacterium_G sp000434315 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Eubacterium_G; Eubacterium_G sp000434315 | |||||||||||
| CAZyme ID | MGYG000000139_01646 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 362709; End: 367604 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 51 | 437 | 2.8e-66 | 0.9662162162162162 |
| GH18 | 661 | 981 | 3.4e-23 | 0.9628378378378378 |
| GH18 | 1333 | 1559 | 1.7e-22 | 0.6587837837837838 |
| CBM5 | 1046 | 1085 | 8e-16 | 0.975 |
| CBM5 | 1235 | 1274 | 8e-16 | 0.975 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06548 | GH18_chitinase | 1.23e-102 | 54 | 429 | 1 | 322 | The GH18 (glycosyl hydrolases, family 18) type II chitinases hydrolyze chitin, an abundant polymer of N-acetylglucosamine and have been identified in bacteria, fungi, insects, plants, viruses, and protozoan parasites. The structure of this domain is an eight-stranded alpha/beta barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. |
| cd02871 | GH18_chitinase_D-like | 2.48e-98 | 1332 | 1617 | 1 | 306 | GH18 domain of Chitinase D (ChiD). ChiD, a chitinase found in Bacillus circulans, hydrolyzes the 1,4-beta-linkages of N-acetylglucosamine in chitin and chitodextrins. The domain architecture of ChiD includes a catalytic glycosyl hydrolase family 18 (GH18) domain, a chitin-binding domain, and a fibronectin type III domain. The chitin-binding and fibronectin type III domains are located either N-terminal or C-terminal to the catalytic domain. This family includes exochitinase Chi36 from Bacillus cereus. |
| cd02871 | GH18_chitinase_D-like | 3.22e-97 | 659 | 975 | 1 | 302 | GH18 domain of Chitinase D (ChiD). ChiD, a chitinase found in Bacillus circulans, hydrolyzes the 1,4-beta-linkages of N-acetylglucosamine in chitin and chitodextrins. The domain architecture of ChiD includes a catalytic glycosyl hydrolase family 18 (GH18) domain, a chitin-binding domain, and a fibronectin type III domain. The chitin-binding and fibronectin type III domains are located either N-terminal or C-terminal to the catalytic domain. This family includes exochitinase Chi36 from Bacillus cereus. |
| COG3325 | ChiA | 3.17e-94 | 35 | 442 | 18 | 436 | Chitinase, GH18 family [Carbohydrate transport and metabolism]. |
| smart00636 | Glyco_18 | 2.36e-85 | 53 | 429 | 1 | 334 | Glyco_18 domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QXF14593.1 | 3.85e-164 | 56 | 593 | 1 | 543 |
| QAV17876.1 | 1.52e-141 | 45 | 990 | 44 | 1084 |
| QXF14594.1 | 2.51e-127 | 120 | 595 | 7 | 472 |
| QOS39875.1 | 2.69e-120 | 656 | 990 | 1047 | 1373 |
| AIY80860.1 | 8.26e-116 | 36 | 579 | 33 | 603 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3EBV_A | 7.35e-90 | 1331 | 1627 | 4 | 294 | Crystalstructure of putative Chitinase A from Streptomyces coelicolor. [Streptomyces coelicolor] |
| 3N11_A | 1.57e-76 | 656 | 991 | 4 | 330 | Crystalstricture of wild-type chitinase from Bacillus cereus NCTU2 [Bacillus cereus],3N12_A Crystal stricture of chitinase in complex with zinc atoms from Bacillus cereus NCTU2 [Bacillus cereus] |
| 3N15_A | 2.91e-76 | 656 | 991 | 4 | 330 | Crystalstricture of E145Q chitinase in complex with NAG from Bacillus cereus NCTU2 [Bacillus cereus] |
| 3N17_A | 9.99e-76 | 656 | 991 | 4 | 330 | Crystalstricture of E145Q/Y227F chitinase in complex with NAG from Bacillus cereus NCTU2 [Bacillus cereus] |
| 3N13_A | 1.85e-75 | 656 | 991 | 4 | 330 | Crystalstricture of D143A chitinase in complex with NAG from Bacillus cereus NCTU2 [Bacillus cereus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P27050 | 9.72e-77 | 658 | 995 | 189 | 519 | Chitinase D OS=Niallia circulans OX=1397 GN=chiD PE=1 SV=4 |
| Q05638 | 1.11e-70 | 661 | 971 | 266 | 581 | Exochitinase 1 OS=Streptomyces olivaceoviridis OX=1921 GN=chi01 PE=1 SV=1 |
| D4AVJ0 | 6.58e-66 | 678 | 990 | 17 | 331 | Probable class II chitinase ARB_00204 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_00204 PE=1 SV=2 |
| P20533 | 3.84e-55 | 23 | 484 | 16 | 495 | Chitinase A1 OS=Niallia circulans OX=1397 GN=chiA1 PE=1 SV=1 |
| E9ERT9 | 5.66e-53 | 52 | 446 | 39 | 403 | Endochitinase 1 OS=Metarhizium robertsii (strain ARSEF 23 / ATCC MYA-3075) OX=655844 GN=chit1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
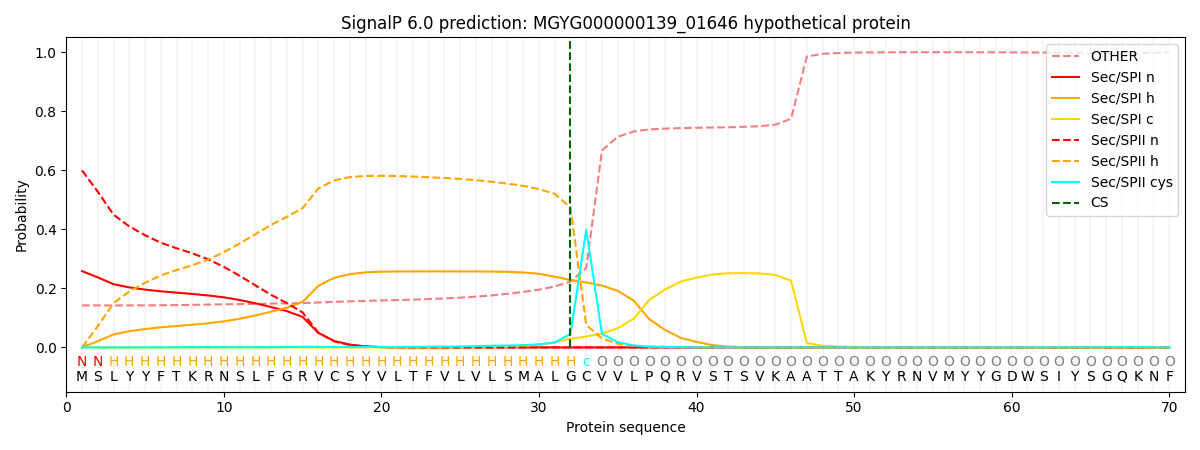
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.148720 | 0.245234 | 0.604003 | 0.000616 | 0.000491 | 0.000925 |