You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000139_02330
You are here: Home > Sequence: MGYG000000139_02330
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
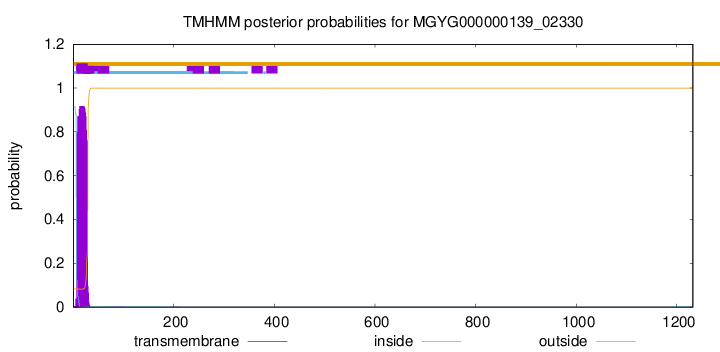
TMHMM annotations
Basic Information help
| Species | Eubacterium_G sp000434315 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Eubacterium_G; Eubacterium_G sp000434315 | |||||||||||
| CAZyme ID | MGYG000000139_02330 | |||||||||||
| CAZy Family | CBM32 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 72518; End: 76216 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CBM32 | 901 | 1015 | 9.2e-20 | 0.9112903225806451 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00754 | F5_F8_type_C | 1.90e-11 | 913 | 1014 | 18 | 127 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| pfam12708 | Pectate_lyase_3 | 7.53e-11 | 54 | 116 | 3 | 66 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| COG5434 | Pgu1 | 2.58e-06 | 26 | 109 | 53 | 138 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| cd00057 | FA58C | 2.10e-05 | 914 | 1003 | 30 | 130 | Substituted updates: Jan 31, 2002 |
| pfam12708 | Pectate_lyase_3 | 1.19e-04 | 485 | 533 | 12 | 61 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AHW60575.1 | 7.83e-288 | 50 | 1017 | 13 | 969 |
| AEV98037.1 | 4.40e-259 | 34 | 1016 | 36 | 1004 |
| QUT90069.1 | 1.85e-233 | 45 | 1015 | 38 | 1030 |
| QJD86468.1 | 3.88e-80 | 34 | 650 | 911 | 1533 |
| QHW29527.1 | 7.38e-64 | 39 | 818 | 1086 | 1964 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7D29_A | 6.96e-06 | 910 | 1024 | 25 | 139 | CBM32of AlyQ [Persicobacter sp. CCB-QB2],7D2A_A CBM32 of AlyQ in complex with 4,5-unsaturated mannuronic acid [Persicobacter sp. CCB-QB2] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
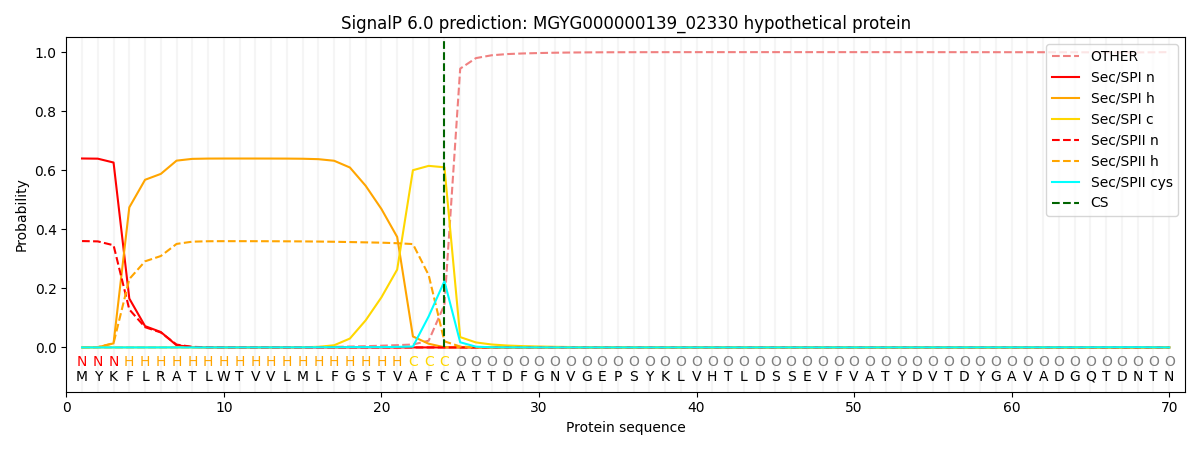
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000927 | 0.632917 | 0.365548 | 0.000212 | 0.000190 | 0.000185 |