You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000150_02160
You are here: Home > Sequence: MGYG000000150_02160
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
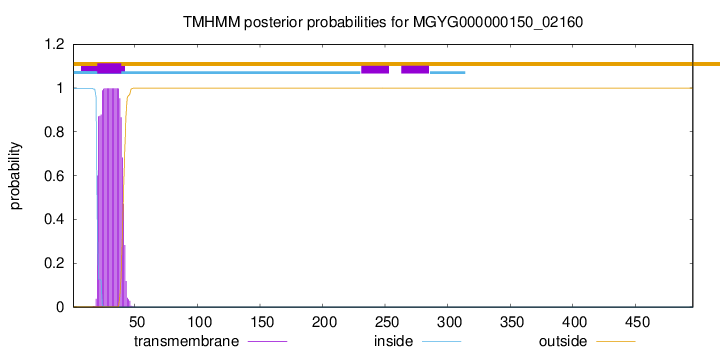
TMHMM annotations
Basic Information help
| Species | Hungatella sp005845265 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Hungatella; Hungatella sp005845265 | |||||||||||
| CAZyme ID | MGYG000000150_02160 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 78494; End: 79984 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE4 | 295 | 410 | 5.2e-29 | 0.8615384615384616 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10944 | CE4_SmPgdA_like | 6.02e-74 | 297 | 484 | 1 | 189 | Catalytic NodB homology domain of Streptococcus mutans polysaccharide deacetylase PgdA, Bacillus subtilis YheN, and similar proteins. This family is represented by a putative polysaccharide deacetylase PgdA from the oral pathogen Streptococcus mutans (SmPgdA) and Bacillus subtilis YheN (BsYheN), which are members of the carbohydrate esterase 4 (CE4) superfamily. SmPgdA is an extracellular metal-dependent polysaccharide deacetylase with a typical CE4 fold, with metal bound to a His-His-Asp triad. It possesses de-N-acetylase activity toward a hexamer of chitooligosaccharide N-acetylglucosamine, but not shorter chitooligosaccharides or a synthetic peptidoglycan tetrasaccharide. SmPgdA plays a role in tuning cell surface properties and in interactions with (salivary) agglutinin, an essential component of the innate immune system, most likely through deacetylation of an as-yet-unidentified polysaccharide. SmPgdA shows significant homology to the catalytic domains of peptidoglycan deacetylases from Streptococcus pneumoniae (SpPgdA) and Listeria monocytogenes (LmPgdA), both of which are involved in the bacterial defense mechanism against human mucosal lysozyme. The Bacillus subtilis genome contains six polysaccharide deacetylase gene homologs: pdaA, pdaB (previously known as ybaN), yheN, yjeA, yxkH and ylxY. The biological function of BsYheN is still unknown. This family also includes many uncharacterized polysaccharide deacetylases mainly found in bacteria. |
| cd02696 | MurNAc-LAA | 9.16e-69 | 120 | 291 | 2 | 172 | N-acetylmuramoyl-L-alanine amidase or MurNAc-LAA (also known as peptidoglycan aminohydrolase, NAMLA amidase, NAMLAA, Amidase 3, and peptidoglycan amidase; EC 3.5.1.28) is an autolysin that hydrolyzes the amide bond between N-acetylmuramoyl and L-amino acids in certain cell wall glycopeptides. These proteins are Zn-dependent peptidases with highly conserved residues involved in cation co-ordination. MurNAc-LAA in this family is one of several peptidoglycan hydrolases (PGHs) found in bacterial and bacteriophage or prophage genomes that are involved in the degradation of the peptidoglycan. In Escherichia coli, there are five MurNAc-LAAs present: AmiA, AmiB, AmiC and AmiD that are periplasmic, and AmpD that is cytoplasmic. Three of these (AmiA, AmiB and AmiC) belong to this family, the other two (AmiD and AmpD) do not. E. coli AmiA, AmiB and AmiC play an important role in cleaving the septum to release daughter cells after cell division. In general, bacterial MurNAc-LAAs are members of the bacterial autolytic system and carry a signal peptide in their N-termini that allows their transport across the cytoplasmic membrane. However, the bacteriophage MurNAc-LAAs are endolysins since these phage-encoded enzymes break down bacterial peptidoglycan at the terminal stage of the phage reproduction cycle. As opposed to autolysins, almost all endolysins have no signal peptides and their translocation through the cytoplasmic membrane is thought to proceed with the help of phage-encoded holin proteins. The amidase catalytic module is fused to another functional module (cell wall binding module or CWBM) either at the N- or C-terminus, which is responsible for high affinity binding of the protein to the cell wall. |
| pfam01520 | Amidase_3 | 7.19e-66 | 120 | 290 | 1 | 172 | N-acetylmuramoyl-L-alanine amidase. This enzyme domain cleaves the amide bond between N-acetylmuramoyl and L-amino acids in bacterial cell walls. |
| COG0860 | AmiC | 4.49e-61 | 117 | 297 | 42 | 231 | N-acetylmuramoyl-L-alanine amidase [Cell wall/membrane/envelope biogenesis]. |
| cd10917 | CE4_NodB_like_6s_7s | 3.33e-49 | 297 | 476 | 1 | 169 | Catalytic NodB homology domain of rhizobial NodB-like proteins. This family belongs to the large and functionally diverse carbohydrate esterase 4 (CE4) superfamily, whose members show strong sequence similarity with some variability due to their distinct carbohydrate substrates. It includes many rhizobial NodB chitooligosaccharide N-deacetylase (EC 3.5.1.-)-like proteins, mainly from bacteria and eukaryotes, such as chitin deacetylases (EC 3.5.1.41), bacterial peptidoglycan N-acetylglucosamine deacetylases (EC 3.5.1.-), and acetylxylan esterases (EC 3.1.1.72), which catalyze the N- or O-deacetylation of substrates such as acetylated chitin, peptidoglycan, and acetylated xylan. All members of this family contain a catalytic NodB homology domain with the same overall topology and a deformed (beta/alpha)8 barrel fold with 6- or 7 strands. Their catalytic activity is dependent on the presence of a divalent cation, preferably cobalt or zinc, and they employ a conserved His-His-Asp zinc-binding triad closely associated with the conserved catalytic base (aspartic acid) and acid (histidine) to carry out acid/base catalysis. Several family members show diversity both in metal ion specificities and in the residues that coordinate the metal. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QJU23357.1 | 3.77e-179 | 118 | 494 | 136 | 512 |
| QRP42504.1 | 3.08e-178 | 106 | 494 | 113 | 502 |
| QOT12681.1 | 7.96e-49 | 108 | 476 | 52 | 461 |
| AWP28024.1 | 2.73e-47 | 109 | 476 | 50 | 459 |
| AYB44783.1 | 7.88e-47 | 108 | 426 | 52 | 412 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4RN7_A | 4.99e-32 | 120 | 297 | 6 | 185 | ChainA, N-acetylmuramoyl-L-alanine amidase [Clostridioides difficile 630] |
| 5EMI_A | 3.19e-27 | 119 | 290 | 6 | 174 | ChainA, Cell wall hydrolase/autolysin [Nostoc punctiforme PCC 73102] |
| 5JMU_A | 2.64e-26 | 297 | 486 | 20 | 214 | ChainA, Peptidoglycan N-acetylglucosamine deacetylase [[Eubacterium] rectale ATCC 33656] |
| 1JWQ_A | 2.26e-23 | 119 | 297 | 3 | 179 | Structureof the catalytic domain of CwlV, N-acetylmuramoyl-L-alanine amidase from Bacillus(Paenibacillus) polymyxa var.colistinus [Paenibacillus polymyxa] |
| 5NC6_A | 4.66e-21 | 297 | 462 | 1 | 164 | Crystalstructure of the polysaccharide deacetylase Bc1974 from Bacillus cereus in complex with (E)-N-hydroxy-3-(naphthalen-1-yl)prop-2-enamide [Bacillus cereus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02114 | 1.71e-33 | 120 | 294 | 322 | 495 | N-acetylmuramoyl-L-alanine amidase LytC OS=Bacillus subtilis (strain 168) OX=224308 GN=lytC PE=1 SV=1 |
| P54525 | 6.52e-33 | 120 | 294 | 32 | 204 | Uncharacterized protein YqiI OS=Bacillus subtilis (strain 168) OX=224308 GN=yqiI PE=3 SV=3 |
| P50864 | 2.48e-29 | 119 | 297 | 42 | 232 | Germination-specific N-acetylmuramoyl-L-alanine amidase OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlD PE=1 SV=1 |
| Q06320 | 1.22e-23 | 120 | 297 | 4 | 178 | Sporulation-specific N-acetylmuramoyl-L-alanine amidase OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlC PE=1 SV=1 |
| P37134 | 2.98e-23 | 120 | 295 | 4 | 176 | N-acetylmuramoyl-L-alanine amidase CwlM OS=Bacillus licheniformis OX=1402 GN=cwlM PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
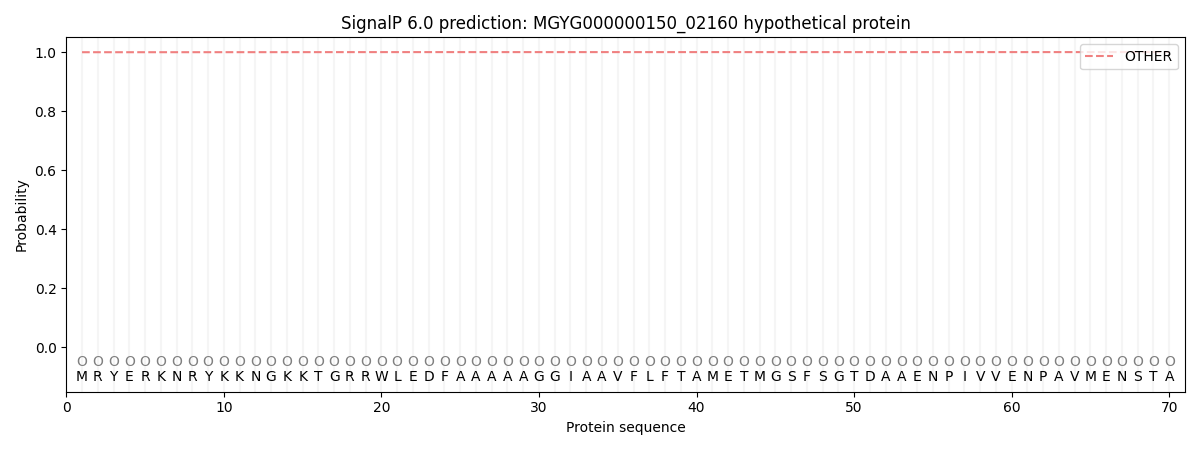
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999801 | 0.000177 | 0.000001 | 0.000001 | 0.000000 | 0.000036 |