You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000151_01994
You are here: Home > Sequence: MGYG000000151_01994
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Clostridium paraputrificum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium; Clostridium paraputrificum | |||||||||||
| CAZyme ID | MGYG000000151_01994 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | Chitodextrinase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 135; End: 794 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06548 | GH18_chitinase | 4.82e-18 | 6 | 60 | 268 | 322 | The GH18 (glycosyl hydrolases, family 18) type II chitinases hydrolyze chitin, an abundant polymer of N-acetylglucosamine and have been identified in bacteria, fungi, insects, plants, viruses, and protozoan parasites. The structure of this domain is an eight-stranded alpha/beta barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. |
| smart00636 | Glyco_18 | 9.06e-16 | 6 | 60 | 280 | 334 | Glyco_18 domain. |
| COG3325 | ChiA | 6.08e-15 | 2 | 61 | 365 | 424 | Chitinase, GH18 family [Carbohydrate transport and metabolism]. |
| pfam06483 | ChiC | 2.51e-14 | 79 | 212 | 2 | 151 | Chitinase C. This ~170 aa region is found at the C-terminus of pfam00704. |
| pfam00704 | Glyco_hydro_18 | 5.56e-13 | 1 | 60 | 250 | 307 | Glycosyl hydrolases family 18. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AWS27147.1 | 1.76e-133 | 1 | 218 | 391 | 608 |
| AEP95044.1 | 1.76e-133 | 1 | 218 | 391 | 608 |
| QIN91090.1 | 1.76e-133 | 1 | 218 | 391 | 608 |
| AEJ34193.1 | 1.76e-133 | 1 | 218 | 391 | 608 |
| AQW28394.1 | 1.76e-133 | 1 | 218 | 391 | 608 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4TXG_A | 1.33e-15 | 18 | 215 | 566 | 774 | CrystalStructure of a Family GH18 Chitinase from Chromobacterium violaceum [Chromobacterium violaceum ATCC 12472] |
| 6BT9_A | 6.16e-12 | 10 | 196 | 398 | 545 | ChitinaseChiA74 from Bacillus thuringiensis [Bacillus thuringiensis],6BT9_B Chitinase ChiA74 from Bacillus thuringiensis [Bacillus thuringiensis] |
| 1ITX_A | 1.42e-10 | 10 | 60 | 356 | 406 | ChainA, Glycosyl Hydrolase [Niallia circulans] |
| 6KST_A | 8.94e-09 | 3 | 60 | 300 | 357 | Crystalstructure of the catalytic domain of chitinase ChiL from Chitiniphilus shinanonensis (CsChiL) [Chitiniphilus shinanonensis],6KST_B Crystal structure of the catalytic domain of chitinase ChiL from Chitiniphilus shinanonensis (CsChiL) [Chitiniphilus shinanonensis],6KXL_A Crystal structure of the catalytic domain of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis],6KXL_B Crystal structure of the catalytic domain of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis] |
| 6KXN_A | 8.94e-09 | 3 | 60 | 300 | 357 | Crystalstructure of W50A mutant of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis],6KXN_B Crystal structure of W50A mutant of Chitiniphilus shinanonensis chitinase ChiL (CsChiL) complexed with N,N'-diacetylchitobiose [Chitiniphilus shinanonensis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P96156 | 9.70e-20 | 11 | 212 | 727 | 955 | Chitodextrinase OS=Vibrio furnissii OX=29494 GN=endo I PE=1 SV=1 |
| A5FB63 | 3.88e-11 | 10 | 200 | 396 | 578 | Chitinase ChiA OS=Flavobacterium johnsoniae (strain ATCC 17061 / DSM 2064 / JCM 8514 / NBRC 14942 / NCIMB 11054 / UW101) OX=376686 GN=chiA PE=1 SV=1 |
| P20533 | 9.35e-10 | 10 | 60 | 388 | 438 | Chitinase A1 OS=Niallia circulans OX=1397 GN=chiA1 PE=1 SV=1 |
| Q9BZP6 | 4.82e-06 | 11 | 122 | 318 | 413 | Acidic mammalian chitinase OS=Homo sapiens OX=9606 GN=CHIA PE=1 SV=1 |
| Q91XA9 | 6.47e-06 | 11 | 94 | 318 | 395 | Acidic mammalian chitinase OS=Mus musculus OX=10090 GN=Chia PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
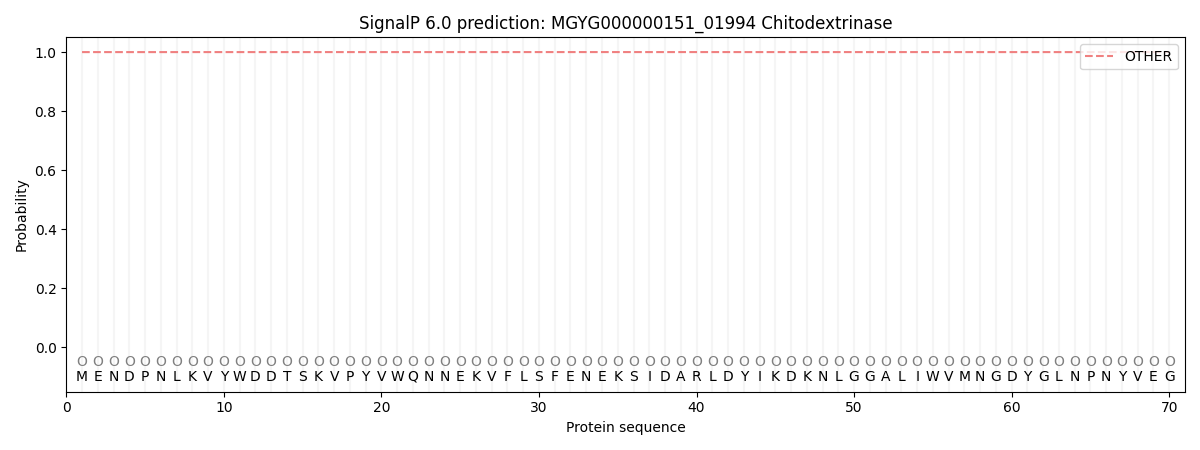
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000052 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
