You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000154_02256
You are here: Home > Sequence: MGYG000000154_02256
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
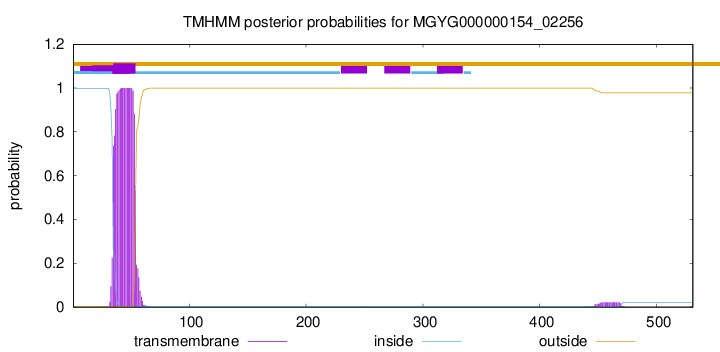
TMHMM annotations
Basic Information help
| Species | RUG115 sp900066395 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; RUG115; RUG115 sp900066395 | |||||||||||
| CAZyme ID | MGYG000000154_02256 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 380882; End: 382477 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01551 | Peptidase_M23 | 3.14e-50 | 431 | 526 | 1 | 96 | Peptidase family M23. Members of this family are zinc metallopeptidases with a range of specificities. The peptidase family M23 is included in this family, these are Gly-Gly endopeptidases. Peptidase family M23 are also endopeptidases. This family also includes some bacterial lipoproteins for which no proteolytic activity has been demonstrated. This family also includes leukocyte cell-derived chemotaxin 2 (LECT2) proteins. LECT2 is a liver-specific protein which is thought to be linked to hepatocyte growth although the exact function of this protein is unknown. |
| COG0739 | NlpD | 1.53e-49 | 276 | 530 | 2 | 261 | Murein DD-endopeptidase MepM and murein hydrolase activator NlpD, contain LysM domain [Cell wall/membrane/envelope biogenesis]. |
| cd12797 | M23_peptidase | 1.54e-41 | 433 | 517 | 1 | 85 | M23 family metallopeptidase, also known as beta-lytic metallopeptidase, and similar proteins. This model describes the metallopeptidase M23 family, which includes beta-lytic metallopeptidase and lysostaphin. Members of this family are zinc endopeptidases that lyse bacterial cell wall peptidoglycans; they cleave either the N-acylmuramoyl-Ala bond between the cell wall peptidoglycan and the cross-linking peptide (e.g. beta-lytic endopeptidase) or a bond within the cross-linking peptide (e.g. stapholysin, and lysostaphin). Beta-lytic metallopeptidase, formerly known as beta-lytic protease, has a preference for cleavage of Gly-X bonds and favors hydrophobic or apolar residues on either side. It inhibits growth of sensitive organisms and may potentially serve as an antimicrobial agent. Lysostaphin, produced by Staphylococcus genus, cleaves pentaglycine cross-bridges of cell wall peptidoglycan, acting as autolysins to maintain cell wall metabolism or as toxins and weapons against competing strains. Staphylolysin (also known as LasA) is implicated in a range of processes related to Pseudomonas virulence, including stimulating shedding of the ectodomain of cell surface heparan sulphate proteoglycan syndecan-1, and elastin degradation in connective tissue. Its active site is less constricted and contains a five-coordinate zinc ion with trigonal bipyramidal geometry and two metal-bound water molecules, possibly contributing to its activity against a wider range of substrates than those used by related lytic enzymes, consistent with its multiple roles in Pseudomonas virulence. The family includes members that do not appear to have the conserved zinc-binding site and might be lipoproteins lacking proteolytic activity. |
| PRK11649 | PRK11649 | 1.54e-33 | 420 | 526 | 293 | 406 | putative peptidase; Provisional |
| PRK10871 | nlpD | 2.81e-20 | 434 | 530 | 220 | 316 | murein hydrolase activator NlpD. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| VCV23561.1 | 1.46e-159 | 30 | 531 | 7 | 519 |
| QNM03019.1 | 1.61e-109 | 59 | 531 | 40 | 528 |
| VDN48046.1 | 3.69e-63 | 204 | 530 | 204 | 541 |
| QUI21292.1 | 3.36e-60 | 214 | 531 | 229 | 551 |
| VDN48047.1 | 2.18e-57 | 219 | 530 | 276 | 594 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3SLU_A | 1.14e-25 | 413 | 528 | 220 | 340 | Crystalstructure of NMB0315 [Neisseria meningitidis ATCC 13091],3SLU_B Crystal structure of NMB0315 [Neisseria meningitidis ATCC 13091] |
| 6MUK_A | 1.72e-25 | 413 | 528 | 240 | 360 | 1.93Angstrom Resolution Crystal Structure of Peptidase M23 from Neisseria gonorrhoeae. [Neisseria gonorrhoeae FA 1090] |
| 5KVP_A | 1.97e-22 | 411 | 518 | 14 | 126 | Solutionstructure of the catalytic domain of zoocin A [Streptococcus equi subsp. zooepidemicus] |
| 6UE4_A | 1.36e-21 | 433 | 525 | 266 | 358 | ShyAEndopeptidase from Vibrio cholerae (Closed form) [Vibrio cholerae O1 biovar El Tor str. N16961],6UE4_B ShyA Endopeptidase from Vibrio cholerae (Closed form) [Vibrio cholerae O1 biovar El Tor str. N16961] |
| 6U2A_A | 1.40e-21 | 433 | 525 | 263 | 355 | ShyAendopeptidase from Vibrio cholera (open form) [Vibrio cholerae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0AFT1 | 1.13e-19 | 420 | 527 | 294 | 408 | Murein DD-endopeptidase MepM OS=Shigella flexneri OX=623 GN=mepM PE=3 SV=1 |
| P0AFS9 | 1.13e-19 | 420 | 527 | 294 | 408 | Murein DD-endopeptidase MepM OS=Escherichia coli (strain K12) OX=83333 GN=mepM PE=1 SV=1 |
| P0AFT0 | 1.13e-19 | 420 | 527 | 294 | 408 | Murein DD-endopeptidase MepM OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=mepM PE=3 SV=1 |
| P44693 | 4.53e-19 | 431 | 527 | 346 | 443 | Uncharacterized metalloprotease HI_0409 OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=HI_0409 PE=3 SV=1 |
| Q8K9M4 | 7.20e-19 | 433 | 527 | 291 | 385 | Uncharacterized metalloprotease BUsg_310 OS=Buchnera aphidicola subsp. Schizaphis graminum (strain Sg) OX=198804 GN=BUsg_310 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
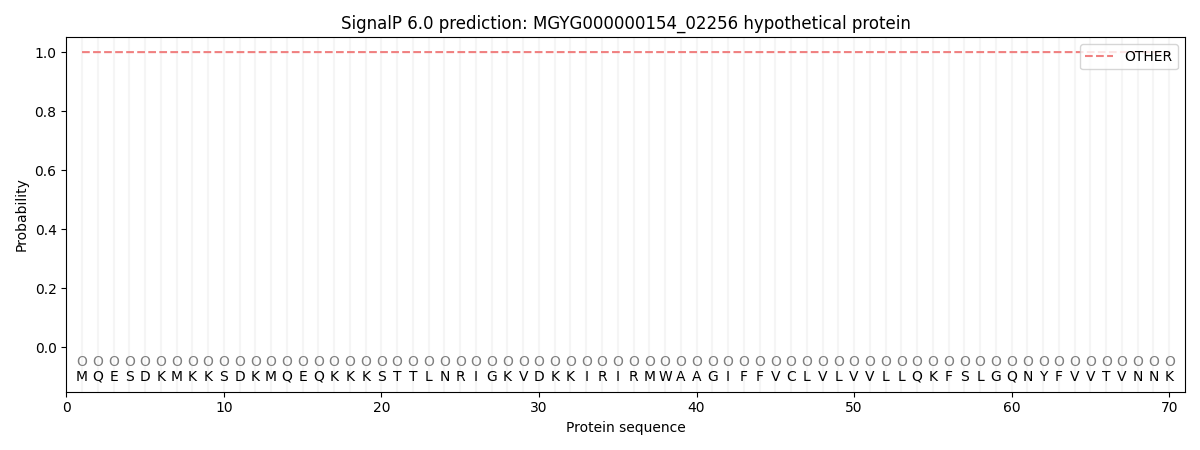
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000001 | 0.000020 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |