You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000157_00343
You are here: Home > Sequence: MGYG000000157_00343
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
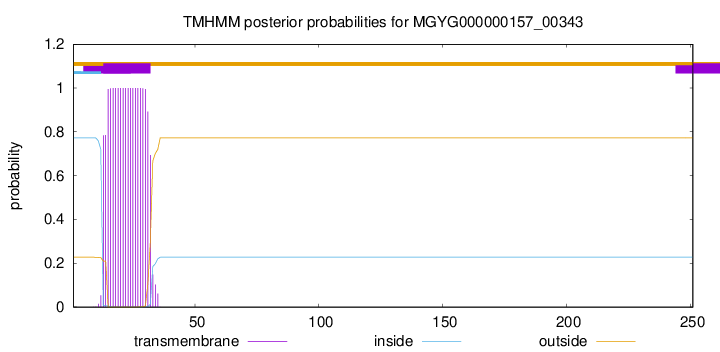
TMHMM annotations
Basic Information help
| Species | Paraclostridium sordellii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Peptostreptococcales; Peptostreptococcaceae; Paraclostridium; Paraclostridium sordellii | |||||||||||
| CAZyme ID | MGYG000000157_00343 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 73856; End: 74611 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10950 | CE4_BsYlxY_like | 2.03e-71 | 55 | 245 | 2 | 188 | Putative catalytic NodB homology domain of uncharacterized protein YlxY from Bacillus subtilis and its bacterial homologs. The Bacillus subtilis genome contains six polysaccharide deacetylase gene homologs: pdaA, pdaB (previously known as ybaN), yheN, yjeA, yxkH and ylxY. This family is represented by Bacillus subtilis putative polysaccharide deacetylase BsYlxY, encoded by the ylxY gene, which is a member of the carbohydrate esterase 4 (CE4) superfamily. Although its biological function still remains unknown, BsYlxY shows high sequence homology to the catalytic domain of Bacillus subtilis pdaB gene encoding a putative polysaccharide deacetylase (BsPdaB), which is essential for the maintenance of spores after the late stage of sporulation and is highly conserved in spore-forming bacteria. However, disruption of the ylxY gene in B. subtilis did not cause any sporulation defect. Moreover, the Asp residue in the classical His-His-Asp zinc-binding motif of CE4 esterases is mutated to a Val residue in this family. Other catalytically relevant residues of CE4 esterases are also not conserved, which suggest that members of this family may be inactive. |
| TIGR02764 | spore_ybaN_pdaB | 3.04e-45 | 56 | 215 | 3 | 158 | polysaccharide deacetylase family sporulation protein PdaB. This model describes the YbaN protein family, also called PdaB and SpoVIE, of Gram-positive bacteria. Although ybaN null mutants have only a mild sporulation defect, ybaN/ytrI double mutants show drastically reducted sporulation efficiencies. This synthetic defect suggests the role of this sigmaE-controlled gene in sporulation had been masked by functional redundancy. Members of this family are homologous to a characterized polysaccharide deacetylase; the exact function this protein family is unknown. [Cellular processes, Sporulation and germination] |
| cd10917 | CE4_NodB_like_6s_7s | 6.79e-45 | 59 | 220 | 1 | 158 | Catalytic NodB homology domain of rhizobial NodB-like proteins. This family belongs to the large and functionally diverse carbohydrate esterase 4 (CE4) superfamily, whose members show strong sequence similarity with some variability due to their distinct carbohydrate substrates. It includes many rhizobial NodB chitooligosaccharide N-deacetylase (EC 3.5.1.-)-like proteins, mainly from bacteria and eukaryotes, such as chitin deacetylases (EC 3.5.1.41), bacterial peptidoglycan N-acetylglucosamine deacetylases (EC 3.5.1.-), and acetylxylan esterases (EC 3.1.1.72), which catalyze the N- or O-deacetylation of substrates such as acetylated chitin, peptidoglycan, and acetylated xylan. All members of this family contain a catalytic NodB homology domain with the same overall topology and a deformed (beta/alpha)8 barrel fold with 6- or 7 strands. Their catalytic activity is dependent on the presence of a divalent cation, preferably cobalt or zinc, and they employ a conserved His-His-Asp zinc-binding triad closely associated with the conserved catalytic base (aspartic acid) and acid (histidine) to carry out acid/base catalysis. Several family members show diversity both in metal ion specificities and in the residues that coordinate the metal. |
| COG0726 | CDA1 | 5.35e-38 | 1 | 218 | 5 | 225 | Peptidoglycan/xylan/chitin deacetylase, PgdA/CDA1 family [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| cd10949 | CE4_BsPdaB_like | 2.56e-32 | 56 | 215 | 1 | 157 | Putative catalytic NodB homology domain of Bacillus subtilis putative polysaccharide deacetylase PdaB, and its bacterial homologs. The Bacillus subtilis genome contains six polysaccharide deacetylase gene homologs: pdaA, pdaB (previously known as ybaN), yheN, yjeA, yxkH and ylxY. This family is represented by the putative polysaccharide deacetylase PdaB encoded by the pdaB gene on sporulation of Bacillus subtilis. Although its biochemical properties remain to be determined, the PdaB (YbaN) protein is essential for maintaining spores after the late stage of sporulation and is highly conserved in spore-forming bacteria. The glycans of the spore cortex may be candidate PdaB substrates. Based on sequence similarity, the family members are classified as carbohydrate esterase 4 (CE4) superfamily members. However, the classical His-His-Asp zinc-binding motif of CE4 esterases is missing in this family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QOX87720.1 | 3.72e-101 | 1 | 245 | 1 | 246 |
| QUP85098.1 | 3.72e-101 | 1 | 245 | 1 | 246 |
| QQY59899.1 | 3.72e-101 | 1 | 245 | 1 | 246 |
| QQY62766.1 | 3.72e-101 | 1 | 245 | 1 | 246 |
| QPK95971.1 | 3.72e-101 | 1 | 245 | 1 | 246 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6HPA_A | 6.66e-39 | 38 | 249 | 66 | 272 | Crystalstructure of a BA3943 mutant,a CE4 family pseudoenzyme [Bacillus anthracis] |
| 6HM9_A | 2.09e-38 | 38 | 249 | 65 | 271 | Crystalstructure of a BA3943 mutant,a CE4 family pseudoenzyme with restored enzymatic activity. [Bacillus anthracis] |
| 7BKF_A | 3.24e-38 | 38 | 249 | 66 | 272 | ChainA, Putative polysaccharide deacetylase [Bacillus anthracis] |
| 2J13_A | 1.41e-19 | 68 | 244 | 62 | 241 | Structureof a family 4 carbohydrate esterase from Bacillus anthracis [Bacillus anthracis str. Ames] |
| 3RXZ_A | 2.84e-08 | 76 | 177 | 64 | 165 | Crystalstructure of putative polysaccharide deacetylase from Mycobacterium smegmatis [Mycolicibacterium smegmatis MC2 155],3RXZ_B Crystal structure of putative polysaccharide deacetylase from Mycobacterium smegmatis [Mycolicibacterium smegmatis MC2 155],3RXZ_C Crystal structure of putative polysaccharide deacetylase from Mycobacterium smegmatis [Mycolicibacterium smegmatis MC2 155],3RXZ_D Crystal structure of putative polysaccharide deacetylase from Mycobacterium smegmatis [Mycolicibacterium smegmatis MC2 155] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P50850 | 8.74e-36 | 37 | 246 | 109 | 313 | Uncharacterized protein YlxY OS=Bacillus subtilis (strain 168) OX=224308 GN=ylxY PE=3 SV=2 |
| P50355 | 6.96e-12 | 55 | 215 | 12 | 171 | Chitooligosaccharide deacetylase OS=Sinorhizobium fredii (strain NBRC 101917 / NGR234) OX=394 GN=nodB PE=3 SV=2 |
| P02963 | 3.51e-11 | 78 | 215 | 38 | 174 | Chitooligosaccharide deacetylase OS=Rhizobium meliloti (strain 1021) OX=266834 GN=nodB PE=3 SV=3 |
| P50352 | 3.61e-11 | 79 | 222 | 39 | 181 | Chitooligosaccharide deacetylase OS=Bradyrhizobium elkanii OX=29448 GN=nodB PE=3 SV=1 |
| P04339 | 1.69e-10 | 78 | 215 | 38 | 174 | Chitooligosaccharide deacetylase OS=Rhizobium leguminosarum bv. viciae OX=387 GN=nodB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.994146 | 0.005500 | 0.000170 | 0.000021 | 0.000013 | 0.000165 |