You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000174_02362
You are here: Home > Sequence: MGYG000000174_02362
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
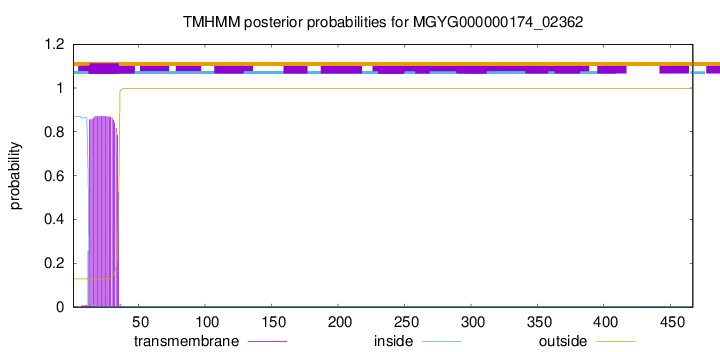
TMHMM annotations
Basic Information help
| Species | Parabacteroides faecis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides faecis | |||||||||||
| CAZyme ID | MGYG000000174_02362 | |||||||||||
| CAZy Family | GH109 | |||||||||||
| CAZyme Description | Alpha-N-acetylgalactosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 348479; End: 349882 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 50 | 460 | 3.1e-170 | 0.9874686716791979 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0673 | MviM | 1.40e-15 | 50 | 415 | 2 | 329 | Predicted dehydrogenase [General function prediction only]. |
| pfam01408 | GFO_IDH_MocA | 9.27e-09 | 52 | 174 | 1 | 113 | Oxidoreductase family, NAD-binding Rossmann fold. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIX66940.1 | 0.0 | 1 | 467 | 1 | 466 |
| QJE27610.1 | 0.0 | 1 | 467 | 1 | 466 |
| ABR42918.1 | 0.0 | 1 | 467 | 1 | 466 |
| QUT18973.1 | 0.0 | 1 | 467 | 1 | 466 |
| QKH97481.1 | 0.0 | 1 | 467 | 1 | 466 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2IXA_A | 1.02e-148 | 51 | 467 | 20 | 440 | A-zyme,N-acetylgalactosaminidase [Elizabethkingia meningoseptica],2IXB_A Crystal structure of N-ACETYLGALACTOSAMINIDASE in complex with GalNAC [Elizabethkingia meningoseptica] |
| 6T2B_A | 5.83e-85 | 50 | 458 | 41 | 438 | Glycosidehydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_B Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_C Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_D Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A6LB54 | 0.0 | 1 | 467 | 1 | 466 | Glycosyl hydrolase family 109 protein OS=Parabacteroides distasonis (strain ATCC 8503 / DSM 20701 / CIP 104284 / JCM 5825 / NCTC 11152) OX=435591 GN=BDI_1155 PE=3 SV=1 |
| A4Q8G1 | 8.64e-301 | 1 | 467 | 1 | 468 | Alpha-N-acetylgalactosaminidase OS=Tannerella forsythia OX=28112 GN=nagA PE=3 SV=1 |
| A4Q8F7 | 5.57e-148 | 51 | 467 | 20 | 440 | Alpha-N-acetylgalactosaminidase OS=Elizabethkingia meningoseptica OX=238 GN=nagA PE=1 SV=1 |
| B2FLK4 | 6.66e-144 | 51 | 467 | 33 | 450 | Glycosyl hydrolase family 109 protein OS=Stenotrophomonas maltophilia (strain K279a) OX=522373 GN=Smlt4431 PE=3 SV=1 |
| A1RI61 | 3.62e-112 | 5 | 463 | 6 | 453 | Glycosyl hydrolase family 109 protein OS=Shewanella sp. (strain W3-18-1) OX=351745 GN=Sputw3181_1518 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as TAT

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 0.000000 | 0.999980 | 0.000000 | 0.000000 |