You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000189_02402
You are here: Home > Sequence: MGYG000000189_02402
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
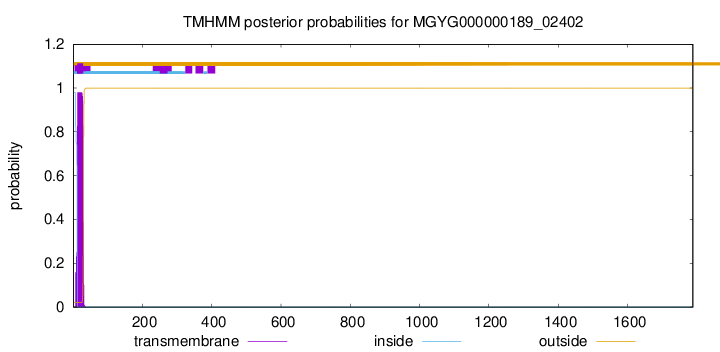
TMHMM annotations
Basic Information help
| Species | Coprococcus sp900066115 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Coprococcus; Coprococcus sp900066115 | |||||||||||
| CAZyme ID | MGYG000000189_02402 | |||||||||||
| CAZy Family | GH9 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 159501; End: 164870 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH9 | 339 | 751 | 3e-102 | 0.9976076555023924 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00759 | Glyco_hydro_9 | 6.62e-105 | 342 | 750 | 1 | 374 | Glycosyl hydrolase family 9. |
| PLN02345 | PLN02345 | 1.08e-48 | 343 | 754 | 1 | 459 | endoglucanase |
| PLN02340 | PLN02340 | 3.38e-46 | 338 | 754 | 29 | 494 | endoglucanase |
| PLN02613 | PLN02613 | 7.57e-43 | 339 | 753 | 26 | 478 | endoglucanase |
| PLN03009 | PLN03009 | 5.52e-41 | 335 | 751 | 24 | 486 | cellulase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK83841.1 | 5.56e-150 | 5 | 1335 | 2 | 1207 |
| AFK82697.1 | 2.56e-132 | 5 | 762 | 2 | 777 |
| ADD61854.1 | 3.69e-125 | 325 | 893 | 179 | 736 |
| QNL98526.1 | 4.93e-125 | 325 | 893 | 198 | 755 |
| QWT52133.1 | 3.96e-120 | 332 | 760 | 30 | 461 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2YIK_A | 4.67e-102 | 337 | 754 | 37 | 513 | ChainA, Endoglucanase [Acetivibrio thermocellus] |
| 1IA6_A | 2.43e-81 | 339 | 752 | 5 | 425 | CrystalStructure Of The Cellulase Cel9m Of C. Cellulolyticum [Ruminiclostridium cellulolyticum],1IA7_A Crystal Structure Of The Cellulase Cel9m Of C. Cellulolyticium In Complex With Cellobiose [Ruminiclostridium cellulolyticum] |
| 2XFG_A | 1.08e-66 | 338 | 757 | 24 | 463 | ChainA, ENDOGLUCANASE 1 [Acetivibrio thermocellus] |
| 4DOD_A | 3.59e-61 | 338 | 757 | 26 | 463 | Thestructure of Cbescii CelA GH9 module [Caldicellulosiruptor bescii],4DOE_A The liganded structure of Cbescii CelA GH9 module [Caldicellulosiruptor bescii] |
| 1JS4_A | 2.95e-59 | 338 | 753 | 4 | 440 | EndoEXOCELLULASE:CELLOBIOSEFROM THERMOMONOSPORA [Thermobifida fusca],1JS4_B EndoEXOCELLULASE:CELLOBIOSE FROM THERMOMONOSPORA [Thermobifida fusca],1TF4_A EndoEXOCELLULASE FROM THERMOMONOSPORA [Thermobifida fusca],1TF4_B EndoEXOCELLULASE FROM THERMOMONOSPORA [Thermobifida fusca],3TF4_A EndoEXOCELLULASE:CELLOTRIOSE FROM THERMOMONOSPORA [Thermobifida fusca],3TF4_B EndoEXOCELLULASE:CELLOTRIOSE FROM THERMOMONOSPORA [Thermobifida fusca],4TF4_A EndoEXOCELLULASE:CELLOPENTAOSE FROM THERMOMONOSPORA [Thermobifida fusca],4TF4_B EndoEXOCELLULASE:CELLOPENTAOSE FROM THERMOMONOSPORA [Thermobifida fusca] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02934 | 1.84e-62 | 338 | 757 | 76 | 515 | Endoglucanase 1 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celI PE=1 SV=2 |
| P22534 | 5.23e-58 | 338 | 757 | 26 | 463 | Endoglucanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=celA PE=3 SV=2 |
| Q5YLG1 | 6.51e-57 | 338 | 751 | 47 | 480 | Endoglucanase A OS=Bacillus pumilus OX=1408 GN=eglA PE=1 SV=1 |
| P26221 | 1.03e-56 | 338 | 753 | 50 | 486 | Endoglucanase E-4 OS=Thermobifida fusca OX=2021 GN=celD PE=1 SV=2 |
| P28622 | 2.78e-55 | 337 | 754 | 27 | 464 | Endoglucanase 4 OS=Bacillus sp. (strain KSM-522) OX=120046 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000414 | 0.998895 | 0.000197 | 0.000162 | 0.000151 | 0.000145 |