You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000193_02369
You are here: Home > Sequence: MGYG000000193_02369
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | KLE1615 sp900066985 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; KLE1615; KLE1615 sp900066985 | |||||||||||
| CAZyme ID | MGYG000000193_02369 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | Xylan 1,3-beta-xylosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 54232; End: 55839 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH43 | 6 | 303 | 1.2e-87 | 0.9928825622775801 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd08989 | GH43_XYL-like | 2.05e-141 | 7 | 298 | 1 | 271 | Glycosyl hydrolase family 43, beta-D-xylosidases and arabinofuranosidases. This glycosyl hydrolase family 43 (GH43) subgroup includes mostly enzymes that have been annotated as having beta-1,4-xylosidase (beta-D-xylosidase;xylan 1,4-beta-xylosidase; EC 3.2.1.37) activity, including Selenomonas ruminantium beta-D-xylosidase SXA. These are part of an array of hemicellulases that are involved in the final breakdown of plant cell-wall whereby they degrade xylan. They hydrolyze beta-1,4 glycosidic bonds between two xylose units in short xylooligosaccharides. It also includes various GH43 family GH43 arabinofuranosidases (EC 3.2.1.55) including Humicola insolens alpha-L-arabinofuranosidase AXHd3, Bacteroides ovatus alpha-L-arabinofuranosidase (BoGH43, XynB), and the bifunctional Phanerochaete chrysosporium xylosidase/arabinofuranosidase (Xyl;PcXyl). GH43 are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Many GH43 enzymes display both alpha-L-arabinofuranosidase and beta-D-xylosidase activity using aryl-glycosides as substrates. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| COG3507 | XynB2 | 2.31e-114 | 1 | 531 | 17 | 549 | Beta-xylosidase [Carbohydrate transport and metabolism]. |
| cd18617 | GH43_XynB-like | 8.03e-108 | 7 | 306 | 1 | 285 | Glycosyl hydrolase family 43, such as Bacteroides ovatus alpha-L-arabinofuranosidase (BoGH43, XynB). This glycosyl hydrolase family 43 (GH43) subgroup includes enzymes that have been characterized to have alpha-L-arabinofuranosidase (EC 3.2.1.55) and beta-1,4-xylosidase (beta-D-xylosidase;xylan 1,4-beta-xylosidase; EC 3.2.1.37) activities. Beta-1,4-xylosidases are part of an array of hemicellulases that are involved in the final breakdown of plant cell-wall whereby they degrade xylan. They hydrolyze beta-1,4 glycosidic bonds between two xylose units in short xylooligosaccharides. These are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Also included in this subfamily are Bacteroides ovatus alpha-L-arabinofuranosidases, BoGH43A and BoGH43B, both having a two-domain architecture, consisting of an N-terminal 5-bladed beta-propeller domain harboring the catalytic active site, and a C-terminal beta-sandwich domain. However, despite significant functional overlap between these two enzymes, BoGH43A and BoGH43B share just 41% sequence identity. The latter appears to be significantly less active on the same substrates, suggesting that these paralogs may play subtly different roles during the degradation of xyloglucans from different sources, or may function most optimally at different stages in the catabolism of xyloglucan oligosaccharides (XyGOs), for example before or after hydrolysis of certain side-chain moieties. It also includes Phanerochaete chrysosporium BKM-F-1767 Xyl, a bifunctional xylosidase/arabinofuranosidase. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| cd09000 | GH43_SXA-like | 1.02e-102 | 7 | 303 | 1 | 289 | Glycosyl hydrolase family 43, such as Selenomonas ruminantium beta-D-xylosidase SXA. This glycosyl hydrolase family 43 (GH43) includes enzymes that have been characterized to mainly have beta-1,4-xylosidase (beta-D-xylosidase;xylan 1,4-beta-xylosidase; EC 3.2.1.37) activity, including Selenomonas ruminantium (Xsa;Sxa;SXA), Bifidobacterium adolescentis ATCC 15703 (XylC;XynB;BAD_0428) and Bacillus sp. KK-1 XylB. They are part of an array of hemicellulases that are involved in the final breakdown of plant cell-wall whereby they degrade xylan. They hydrolyze beta-1,4 glycosidic bonds between two xylose units in short xylooligosaccharides. These are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. These enzymes possess an additional C-terminal beta-sandwich domain that restricts access for substrates to a portion of the active site to form a pocket. The active-site pockets comprise of two subsites, with binding capacity for two monosaccharide moieties and a single route of access for small molecules such as substrate. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| pfam04616 | Glyco_hydro_43 | 8.39e-88 | 5 | 304 | 1 | 281 | Glycosyl hydrolases family 43. The glycosyl hydrolase family 43 contains members that are arabinanases. Arabinanases hydrolyze the alpha-1,5-linked L-arabinofuranoside backbone of plant cell wall arabinans. The structure of arabinanase Arb43A from Cellvibrio japonicus reveals a five-bladed beta-propeller fold. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACR71981.1 | 1.53e-310 | 1 | 529 | 1 | 511 |
| ABX40578.1 | 4.68e-291 | 1 | 529 | 1 | 511 |
| BCJ93959.1 | 5.15e-290 | 3 | 529 | 11 | 519 |
| QNM01562.1 | 9.66e-289 | 3 | 529 | 5 | 513 |
| AUO19465.1 | 1.32e-226 | 5 | 529 | 3 | 509 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6MS2_A | 9.86e-220 | 4 | 528 | 29 | 535 | Crystalstructure of the GH43 BlXynB protein from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580] |
| 6MS3_A | 5.66e-219 | 4 | 528 | 29 | 535 | Crystalstructure of the GH43 protein BlXynB mutant (K247S) from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580],6MS3_B Crystal structure of the GH43 protein BlXynB mutant (K247S) from Bacillus licheniformis [Bacillus licheniformis DSM 13 = ATCC 14580] |
| 1YRZ_A | 3.25e-108 | 1 | 532 | 1 | 527 | ChainA, xylan beta-1,4-xylosidase [Halalkalibacterium halodurans C-125],1YRZ_B Chain B, xylan beta-1,4-xylosidase [Halalkalibacterium halodurans C-125] |
| 3C2U_A | 9.84e-96 | 6 | 532 | 4 | 538 | Structureof the two subsite D-xylosidase from Selenomonas ruminantium in complex with 1,3-bis[tris(hydroxymethyl)methylamino]propane [Selenomonas ruminantium],3C2U_B Structure of the two subsite D-xylosidase from Selenomonas ruminantium in complex with 1,3-bis[tris(hydroxymethyl)methylamino]propane [Selenomonas ruminantium],3C2U_C Structure of the two subsite D-xylosidase from Selenomonas ruminantium in complex with 1,3-bis[tris(hydroxymethyl)methylamino]propane [Selenomonas ruminantium],3C2U_D Structure of the two subsite D-xylosidase from Selenomonas ruminantium in complex with 1,3-bis[tris(hydroxymethyl)methylamino]propane [Selenomonas ruminantium] |
| 2EXH_A | 9.91e-95 | 6 | 529 | 5 | 533 | Structureof the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus],2EXH_B Structure of the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus],2EXH_C Structure of the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus],2EXH_D Structure of the family43 beta-Xylosidase from geobacillus stearothermophilus [Geobacillus stearothermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A9ZND1 | 1.90e-98 | 1 | 528 | 1 | 532 | Xylan 1,3-beta-xylosidase OS=Vibrio sp. OX=678 GN=xloA PE=1 SV=1 |
| A7LXT8 | 1.29e-92 | 5 | 531 | 23 | 521 | Non-reducing end alpha-L-arabinofuranosidase BoGH43A OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02654 PE=1 SV=1 |
| P77713 | 1.43e-90 | 7 | 529 | 5 | 534 | Putative beta-xylosidase OS=Escherichia coli (strain K12) OX=83333 GN=yagH PE=3 SV=1 |
| A7LXU0 | 1.40e-88 | 3 | 532 | 25 | 529 | Non-reducing end alpha-L-arabinofuranosidase BoGH43B OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02656 PE=1 SV=2 |
| P94489 | 7.16e-86 | 7 | 529 | 5 | 531 | Beta-xylosidase OS=Bacillus subtilis (strain 168) OX=224308 GN=xynB PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
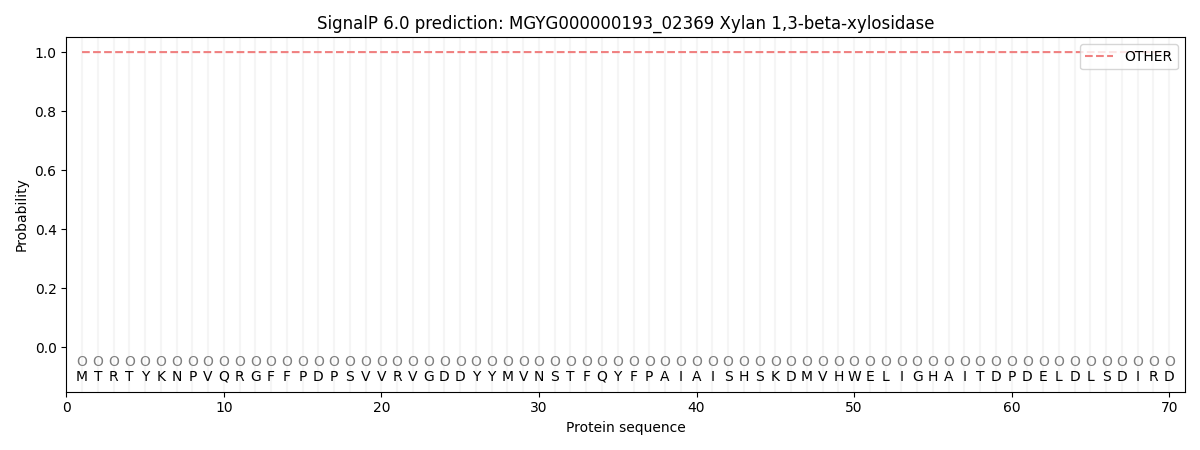
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000036 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
