You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000193_02482
You are here: Home > Sequence: MGYG000000193_02482
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | KLE1615 sp900066985 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; KLE1615; KLE1615 sp900066985 | |||||||||||
| CAZyme ID | MGYG000000193_02482 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-glucuronidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 47713; End: 49494 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 9 | 588 | 1.7e-101 | 0.6090425531914894 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10150 | PRK10150 | 0.0 | 1 | 590 | 1 | 592 | beta-D-glucuronidase; Provisional |
| COG3250 | LacZ | 2.08e-108 | 1 | 590 | 1 | 595 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| pfam02836 | Glyco_hydro_2_C | 1.02e-90 | 285 | 590 | 1 | 297 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| PRK10340 | ebgA | 3.69e-36 | 10 | 508 | 39 | 514 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam02837 | Glyco_hydro_2_N | 1.89e-29 | 12 | 186 | 1 | 169 | Glycosyl hydrolases family 2, sugar binding domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities and has a jelly-roll fold. The domain binds the sugar moiety during the sugar-hydrolysis reaction. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFH61161.1 | 2.69e-245 | 1 | 590 | 1 | 589 |
| AFC28929.1 | 5.40e-245 | 1 | 590 | 1 | 589 |
| AEI40295.1 | 7.66e-245 | 1 | 590 | 1 | 589 |
| AZT89837.1 | 1.45e-227 | 1 | 590 | 1 | 588 |
| ADL43136.1 | 2.63e-226 | 1 | 590 | 1 | 588 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6U7J_A | 1.89e-315 | 1 | 590 | 10 | 590 | UnculturedClostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_B Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_C Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_D Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.] |
| 4JKM_A | 1.33e-203 | 1 | 590 | 4 | 592 | CrystalStructure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],4JKM_B Crystal Structure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],6CXS_A Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13],6CXS_B Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13] |
| 6XXW_A | 4.02e-199 | 1 | 590 | 21 | 589 | Structureof beta-D-Glucuronidase for Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12] |
| 5C70_A | 3.14e-194 | 1 | 590 | 9 | 597 | Thestructure of Aspergillus oryzae beta-glucuronidase [Aspergillus oryzae],5C70_B The structure of Aspergillus oryzae beta-glucuronidase [Aspergillus oryzae] |
| 5C71_A | 5.84e-193 | 1 | 590 | 34 | 622 | Thestructure of Aspergillus oryzae a-glucuronidase complexed with glycyrrhetinic acid monoglucuronide [Aspergillus oryzae],5C71_B The structure of Aspergillus oryzae a-glucuronidase complexed with glycyrrhetinic acid monoglucuronide [Aspergillus oryzae],5C71_C The structure of Aspergillus oryzae a-glucuronidase complexed with glycyrrhetinic acid monoglucuronide [Aspergillus oryzae],5C71_D The structure of Aspergillus oryzae a-glucuronidase complexed with glycyrrhetinic acid monoglucuronide [Aspergillus oryzae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P12265 | 1.00e-193 | 1 | 590 | 27 | 622 | Beta-glucuronidase OS=Mus musculus OX=10090 GN=Gusb PE=1 SV=2 |
| O97524 | 1.79e-192 | 1 | 590 | 27 | 625 | Beta-glucuronidase OS=Felis catus OX=9685 GN=GUSB PE=1 SV=1 |
| O18835 | 7.19e-192 | 1 | 590 | 27 | 625 | Beta-glucuronidase OS=Canis lupus familiaris OX=9615 GN=GUSB PE=1 SV=1 |
| Q4FAT7 | 9.66e-190 | 1 | 590 | 28 | 626 | Beta-glucuronidase OS=Sus scrofa OX=9823 GN=GUSB PE=3 SV=1 |
| P08236 | 1.87e-189 | 1 | 590 | 27 | 626 | Beta-glucuronidase OS=Homo sapiens OX=9606 GN=GUSB PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
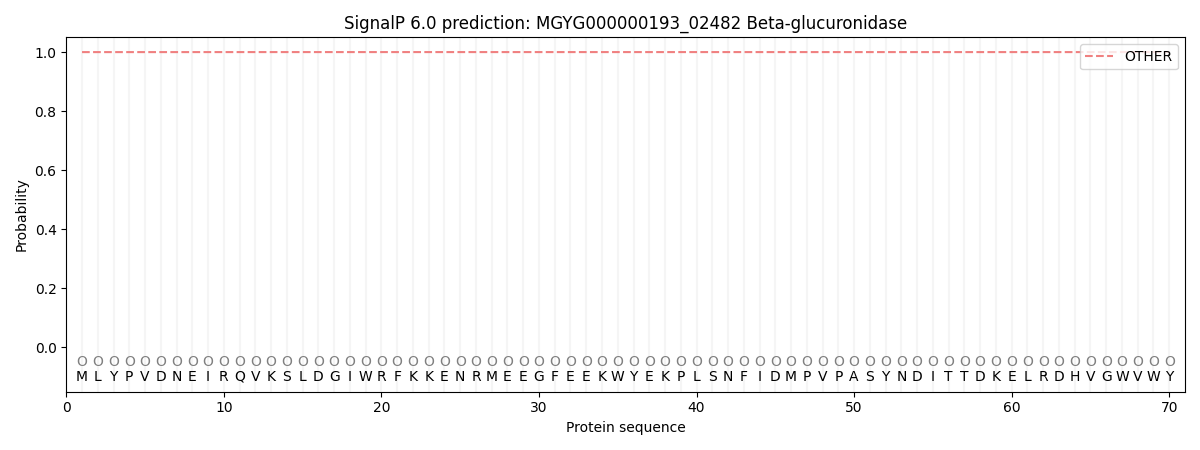
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000042 | 0.000002 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
