You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000196_01750
You are here: Home > Sequence: MGYG000000196_01750
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides thetaiotaomicron | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides thetaiotaomicron | |||||||||||
| CAZyme ID | MGYG000000196_01750 | |||||||||||
| CAZy Family | CE8 | |||||||||||
| CAZyme Description | Pectinesterase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 229242; End: 230210 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE8 | 32 | 313 | 1.4e-104 | 0.9895833333333334 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PLN02773 | PLN02773 | 1.48e-80 | 26 | 306 | 1 | 290 | pectinesterase |
| pfam01095 | Pectinesterase | 9.47e-79 | 32 | 310 | 2 | 293 | Pectinesterase. |
| PLN02432 | PLN02432 | 2.34e-73 | 32 | 306 | 13 | 280 | putative pectinesterase |
| COG4677 | PemB | 3.13e-70 | 17 | 313 | 72 | 403 | Pectin methylesterase and related acyl-CoA thioesterases [Carbohydrate transport and metabolism, Lipid transport and metabolism]. |
| PLN02682 | PLN02682 | 3.73e-68 | 31 | 303 | 70 | 352 | pectinesterase family protein |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CAW41305.1 | 3.26e-226 | 1 | 322 | 1 | 322 |
| ALJ43642.1 | 3.26e-226 | 1 | 322 | 1 | 322 |
| CBC06956.1 | 3.26e-226 | 1 | 322 | 1 | 322 |
| QUT69789.1 | 3.26e-226 | 1 | 322 | 1 | 322 |
| AAO79214.1 | 3.26e-226 | 1 | 322 | 1 | 322 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1GQ8_A | 8.21e-34 | 32 | 308 | 9 | 298 | Pectinmethylesterase from Carrot [Daucus carota] |
| 1XG2_A | 1.10e-33 | 33 | 313 | 6 | 296 | ChainA, Pectinesterase 1 [Solanum lycopersicum] |
| 5C1E_A | 1.61e-26 | 32 | 291 | 11 | 271 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Penultimately Deglycosylated Form (N-acetylglucosamine Stub at Asn84) [Aspergillus niger ATCC 1015] |
| 5C1C_A | 8.28e-26 | 32 | 291 | 11 | 271 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Deglycosylated Form [Aspergillus niger ATCC 1015] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9LVQ0 | 5.68e-50 | 34 | 306 | 9 | 290 | Pectinesterase 31 OS=Arabidopsis thaliana OX=3702 GN=PME31 PE=1 SV=1 |
| P41510 | 1.18e-45 | 31 | 316 | 272 | 577 | Probable pectinesterase/pectinesterase inhibitor OS=Brassica napus OX=3708 GN=BP19 PE=2 SV=1 |
| Q42608 | 3.63e-44 | 31 | 316 | 259 | 564 | Pectinesterase/pectinesterase inhibitor (Fragment) OS=Brassica campestris OX=3711 PE=2 SV=1 |
| Q9LXD9 | 1.02e-42 | 29 | 318 | 238 | 546 | Probable pectinesterase/pectinesterase inhibitor 51 OS=Arabidopsis thaliana OX=3702 GN=PME51 PE=2 SV=1 |
| Q8VYZ3 | 2.24e-42 | 33 | 318 | 87 | 380 | Probable pectinesterase 53 OS=Arabidopsis thaliana OX=3702 GN=PME53 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
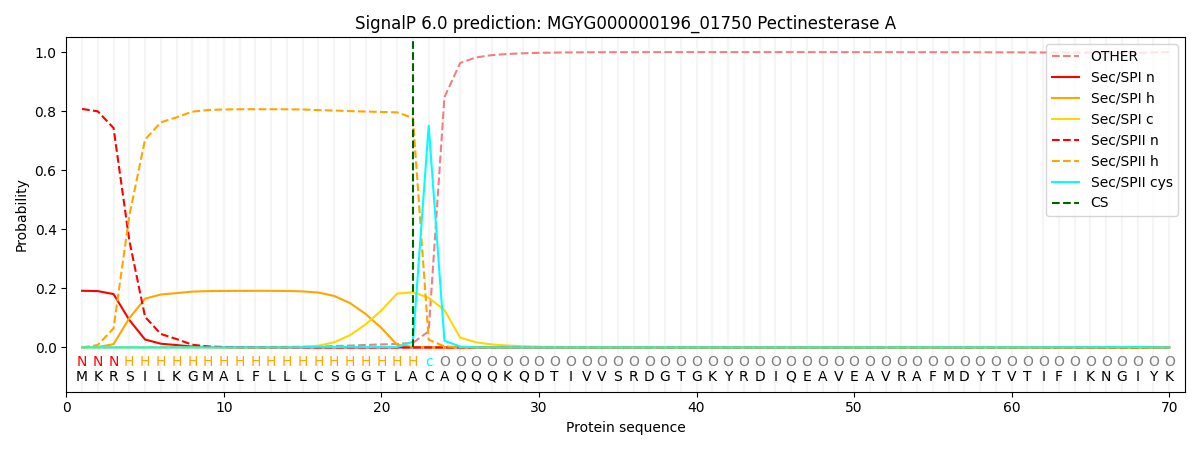
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001994 | 0.184717 | 0.812634 | 0.000219 | 0.000220 | 0.000212 |
