You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000198_01888
You are here: Home > Sequence: MGYG000000198_01888
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Enterocloster citroniae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Enterocloster; Enterocloster citroniae | |||||||||||
| CAZyme ID | MGYG000000198_01888 | |||||||||||
| CAZy Family | CE15 | |||||||||||
| CAZyme Description | Carbohydrate esterase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 768637; End: 769764 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE15 | 11 | 374 | 1.3e-85 | 0.9851301115241635 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1506 | DAP2 | 2.58e-05 | 195 | 239 | 462 | 506 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| pfam00326 | Peptidase_S9 | 0.002 | 195 | 237 | 53 | 96 | Prolyl oligopeptidase family. |
| pfam00561 | Abhydrolase_1 | 0.003 | 188 | 326 | 53 | 178 | alpha/beta hydrolase fold. This catalytic domain is found in a very wide range of enzymes. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QSI03296.1 | 9.13e-170 | 9 | 373 | 8 | 374 |
| AEV29352.1 | 1.32e-155 | 29 | 374 | 24 | 369 |
| QHW35286.1 | 5.91e-119 | 7 | 374 | 3 | 367 |
| QHT60959.1 | 6.26e-117 | 7 | 373 | 3 | 366 |
| QNK58798.1 | 2.30e-112 | 4 | 372 | 1 | 369 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6EHN_A | 1.95e-79 | 8 | 374 | 11 | 399 | Structuralinsight into a promiscuous CE15 esterase from the marine bacterial metagenome [unidentified prokaryotic organism] |
| 6GRW_A | 3.83e-77 | 11 | 375 | 7 | 398 | GlucuronoylEsterase from Opitutus terrae (Au derivative) [Opitutus terrae PB90-1],6GS0_A Native Glucuronoyl Esterase from Opitutus terrae [Opitutus terrae PB90-1] |
| 6SYU_A | 6.33e-77 | 11 | 375 | 25 | 416 | Thewild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with xylobiose [Opitutus terrae],6T0I_A The wild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with the aldotetrauronic acid XUX [Opitutus terrae] |
| 6SYR_A | 6.96e-77 | 5 | 375 | 30 | 427 | Thewild type glucuronoyl esterase OtCE15A from Opitutus terrae in complex with D-glucuronate [Opitutus terrae PB90-1] |
| 6GRY_A | 9.07e-77 | 11 | 370 | 14 | 392 | GlucuronoylEsterase from Solibacter usitatus. [Candidatus Solibacter usitatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0A0K2VM55 | 2.71e-78 | 8 | 374 | 39 | 427 | Carbohydrate esterase MZ0003 OS=Unknown prokaryotic organism OX=2725 GN=MZ0003 PE=1 SV=1 |
| D8QLP9 | 2.33e-21 | 11 | 328 | 36 | 326 | 4-O-methyl-glucuronoyl methylesterase OS=Schizophyllum commune (strain H4-8 / FGSC 9210) OX=578458 GN=SCHCODRAFT_238770 PE=1 SV=1 |
| A0A0A7EQR3 | 4.27e-21 | 7 | 327 | 108 | 406 | 4-O-methyl-glucuronoyl methylesterase OS=Cerrena unicolor OX=90312 PE=1 SV=1 |
| K5XDZ6 | 4.16e-20 | 4 | 328 | 37 | 338 | 4-O-methyl-glucuronoyl methylesterase OS=Phanerochaete carnosa (strain HHB-10118-sp) OX=650164 GN=PHACADRAFT_247750 PE=1 SV=1 |
| P0CU53 | 1.43e-19 | 7 | 328 | 41 | 340 | 4-O-methyl-glucuronoyl methylesterase 1 OS=Wolfiporia cocos (strain MD-104) OX=742152 GN=WOLCODRAFT_23632 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
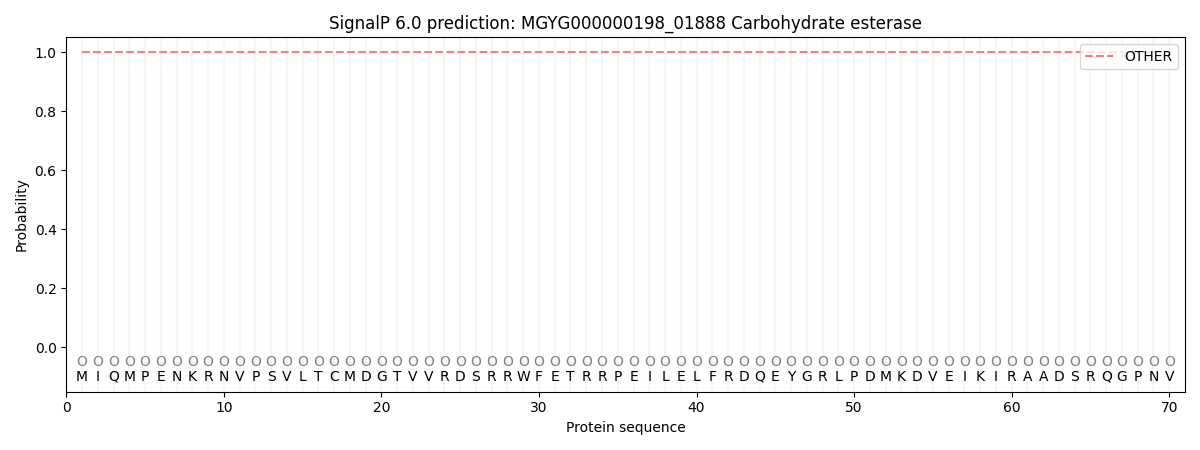
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000059 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
