You are browsing environment: HUMAN GUT
MGYG000000200_00059
Basic Information
help
Species
Blautia_A sp003471165
Lineage
Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Blautia_A; Blautia_A sp003471165
CAZyme ID
MGYG000000200_00059
CAZy Family
GH73
CAZyme Description
hypothetical protein
CAZyme Property
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000000200
4192583
Isolate
China
Asia
Gene Location
Start: 64988;
End: 65743
Strand: -
No EC number prediction in MGYG000000200_00059.
Family
Start
End
Evalue
family coverage
GH73
119
248
3.5e-18
0.9609375
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
pfam01832
Glucosaminidase
1.37e-09
119
198
1
91
Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase. This family includes Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase EC:3.2.1.96. As well as the flageller protein J that has been shown to hydrolyze peptidoglycan.
more
Hit ID
E-Value
Query Start
Query End
Hit Start
Hit End
Description
O51481
5.97e-22
115
249
68
195
Uncharacterized protein BB_0531 OS=Borreliella burgdorferi (strain ATCC 35210 / DSM 4680 / CIP 102532 / B31) OX=224326 GN=BB_0531 PE=3 SV=1
more
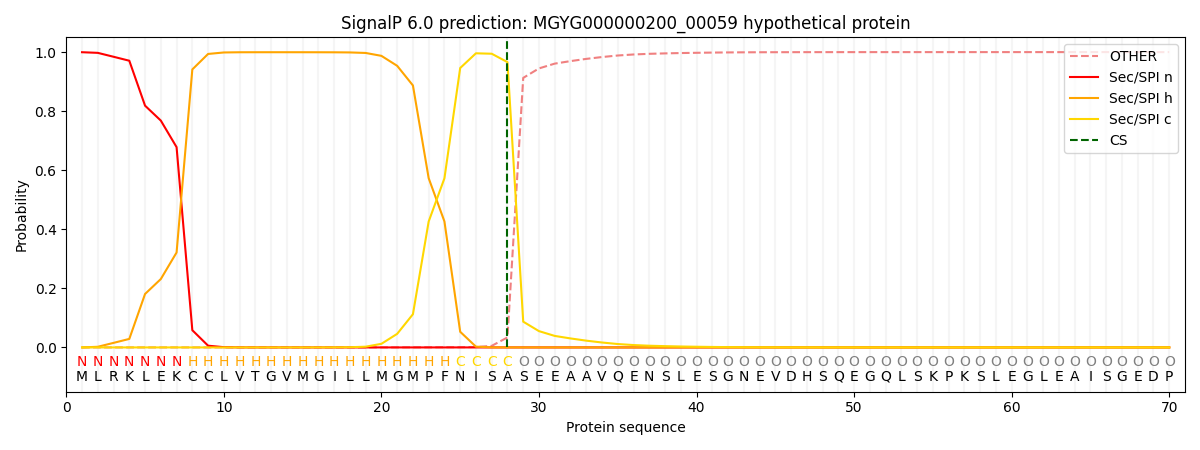
This protein is predicted as SP
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
0.000616
0.998007
0.000781
0.000182
0.000191
0.000176
There is no transmembrane helices in MGYG000000200_00059.