You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000202_03920
You are here: Home > Sequence: MGYG000000202_03920
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | AF33-28 sp003477885 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; AF33-28; AF33-28 sp003477885 | |||||||||||
| CAZyme ID | MGYG000000202_03920 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 40602; End: 43940 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 2 | 478 | 3e-57 | 0.48404255319148937 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 1.75e-15 | 2 | 526 | 13 | 503 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10150 | PRK10150 | 1.19e-14 | 3 | 333 | 14 | 298 | beta-D-glucuronidase; Provisional |
| pfam00754 | F5_F8_type_C | 3.85e-10 | 951 | 1047 | 15 | 114 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| PRK10340 | ebgA | 3.10e-08 | 4 | 433 | 44 | 451 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam02836 | Glyco_hydro_2_C | 1.99e-06 | 311 | 483 | 1 | 185 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ABX43438.1 | 0.0 | 1 | 1111 | 1 | 1148 |
| BCJ95530.1 | 0.0 | 4 | 1111 | 9 | 1148 |
| QJE01939.1 | 2.08e-306 | 2 | 1055 | 17 | 1071 |
| ARN57843.1 | 1.26e-305 | 2 | 1049 | 33 | 1057 |
| QGZ42623.1 | 6.02e-305 | 2 | 1055 | 44 | 1068 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1JZ7_A | 7.57e-14 | 4 | 445 | 54 | 476 | E.COLI (lacZ) BETA-GALACTOSIDASE IN COMPLEX WITH GALACTOSE [Escherichia coli],1JZ7_B E. COLI (lacZ) BETA-GALACTOSIDASE IN COMPLEX WITH GALACTOSE [Escherichia coli],1JZ7_C E. COLI (lacZ) BETA-GALACTOSIDASE IN COMPLEX WITH GALACTOSE [Escherichia coli],1JZ7_D E. COLI (lacZ) BETA-GALACTOSIDASE IN COMPLEX WITH GALACTOSE [Escherichia coli],4TTG_A Beta-galactosidase (E. coli) in the presence of potassium chloride. [Escherichia coli],4TTG_B Beta-galactosidase (E. coli) in the presence of potassium chloride. [Escherichia coli],4TTG_C Beta-galactosidase (E. coli) in the presence of potassium chloride. [Escherichia coli],4TTG_D Beta-galactosidase (E. coli) in the presence of potassium chloride. [Escherichia coli] |
| 4JKM_A | 1.17e-13 | 1 | 446 | 15 | 436 | CrystalStructure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],4JKM_B Crystal Structure of Clostridium perfringens beta-glucuronidase [Clostridium perfringens str. 13],6CXS_A Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13],6CXS_B Crystal Structure of Clostridium perfringens beta-glucuronidase bound with a novel, potent inhibitor 4-(8-(piperazin-1-yl)-1,2,3,4-tetrahydro-[1,2,3]triazino[4',5':4,5]thieno[2,3-c]isoquinolin-5-yl)morpholine [Clostridium perfringens str. 13] |
| 3MUY_1 | 1.30e-13 | 4 | 445 | 54 | 476 | Chain1, Beta-D-galactosidase [Escherichia coli K-12],3MUY_2 Chain 2, Beta-D-galactosidase [Escherichia coli K-12],3MUY_3 Chain 3, Beta-D-galactosidase [Escherichia coli K-12],3MUY_4 Chain 4, Beta-D-galactosidase [Escherichia coli K-12] |
| 3IAQ_A | 1.30e-13 | 4 | 445 | 54 | 476 | ChainA, Beta-galactosidase [Escherichia coli K-12],3IAQ_B Chain B, Beta-galactosidase [Escherichia coli K-12],3IAQ_C Chain C, Beta-galactosidase [Escherichia coli K-12],3IAQ_D Chain D, Beta-galactosidase [Escherichia coli K-12] |
| 3J7H_A | 1.30e-13 | 4 | 445 | 55 | 477 | Structureof beta-galactosidase at 3.2-A resolution obtained by cryo-electron microscopy [Escherichia coli K-12],3J7H_B Structure of beta-galactosidase at 3.2-A resolution obtained by cryo-electron microscopy [Escherichia coli K-12],3J7H_C Structure of beta-galactosidase at 3.2-A resolution obtained by cryo-electron microscopy [Escherichia coli K-12],3J7H_D Structure of beta-galactosidase at 3.2-A resolution obtained by cryo-electron microscopy [Escherichia coli K-12],4CKD_A Model of complex between the E.coli enzyme beta-galactosidase and four single chain Fv antibody domains scFv13R4. [Escherichia coli K-12],4CKD_B Model of complex between the E.coli enzyme beta-galactosidase and four single chain Fv antibody domains scFv13R4. [Escherichia coli K-12],4CKD_C Model of complex between the E.coli enzyme beta-galactosidase and four single chain Fv antibody domains scFv13R4. [Escherichia coli K-12],4CKD_D Model of complex between the E.coli enzyme beta-galactosidase and four single chain Fv antibody domains scFv13R4. [Escherichia coli K-12],6DRV_A Beta-galactosidase [Escherichia coli K-12],6DRV_B Beta-galactosidase [Escherichia coli K-12],6DRV_C Beta-galactosidase [Escherichia coli K-12],6DRV_D Beta-galactosidase [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B7N8Q1 | 3.16e-13 | 4 | 445 | 55 | 477 | Beta-galactosidase OS=Escherichia coli O17:K52:H18 (strain UMN026 / ExPEC) OX=585056 GN=lacZ PE=3 SV=1 |
| A9MQ82 | 5.44e-13 | 4 | 445 | 55 | 477 | Beta-galactosidase OS=Salmonella arizonae (strain ATCC BAA-731 / CDC346-86 / RSK2980) OX=41514 GN=lacZ PE=3 SV=2 |
| Q32JB6 | 5.44e-13 | 4 | 445 | 55 | 477 | Beta-galactosidase OS=Shigella dysenteriae serotype 1 (strain Sd197) OX=300267 GN=lacZ PE=3 SV=2 |
| Q0TKT1 | 7.14e-13 | 4 | 445 | 55 | 477 | Beta-galactosidase OS=Escherichia coli O6:K15:H31 (strain 536 / UPEC) OX=362663 GN=lacZ PE=3 SV=1 |
| Q1RFJ2 | 7.14e-13 | 4 | 445 | 55 | 477 | Beta-galactosidase OS=Escherichia coli (strain UTI89 / UPEC) OX=364106 GN=lacZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
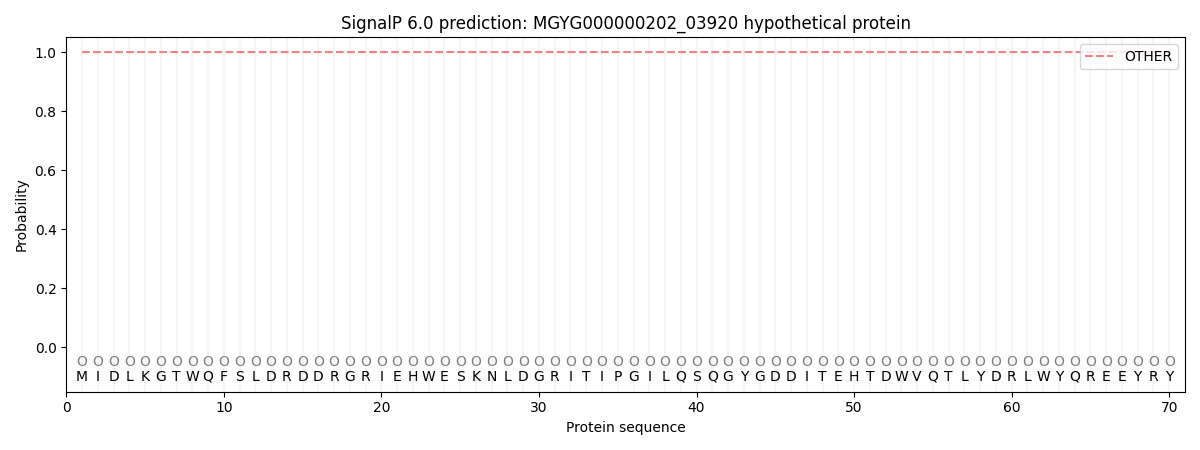
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000046 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
