You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000222_00790
You are here: Home > Sequence: MGYG000000222_00790
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides sp003473295 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides sp003473295 | |||||||||||
| CAZyme ID | MGYG000000222_00790 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 92437; End: 93873 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH30 | 33 | 473 | 3.1e-140 | 0.9978070175438597 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam14587 | Glyco_hydr_30_2 | 5.84e-26 | 33 | 346 | 4 | 353 | O-Glycosyl hydrolase family 30. |
| COG5520 | XynC | 2.72e-05 | 142 | 477 | 114 | 430 | O-Glycosyl hydrolase [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBE17217.1 | 7.08e-228 | 33 | 477 | 32 | 478 |
| AEW47939.1 | 1.54e-204 | 35 | 477 | 39 | 485 |
| AEW47948.1 | 1.54e-204 | 35 | 477 | 39 | 485 |
| ADF52950.1 | 1.58e-188 | 35 | 474 | 28 | 468 |
| AUX29323.1 | 5.17e-107 | 31 | 477 | 84 | 523 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5CXP_A | 2.13e-07 | 300 | 476 | 218 | 388 | X-raycrystallographic protein structure of the glycoside hydrolase family 30 subfamily 8 xylanase, Xyn30A, from Clostridium acetobutylicum [Clostridium acetobutylicum ATCC 824] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q76FP5 | 1.47e-49 | 30 | 415 | 21 | 409 | Endo-beta-1,6-galactanase OS=Hypocrea rufa OX=5547 GN=6GAL PE=1 SV=1 |
| A0A401ETL2 | 1.18e-20 | 33 | 351 | 37 | 446 | Exo-beta-1,6-galactobiohydrolase OS=Bifidobacterium longum subsp. longum (strain ATCC 15707 / DSM 20219 / JCM 1217 / NCTC 11818 / E194b) OX=565042 GN=bl1,6Gal PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
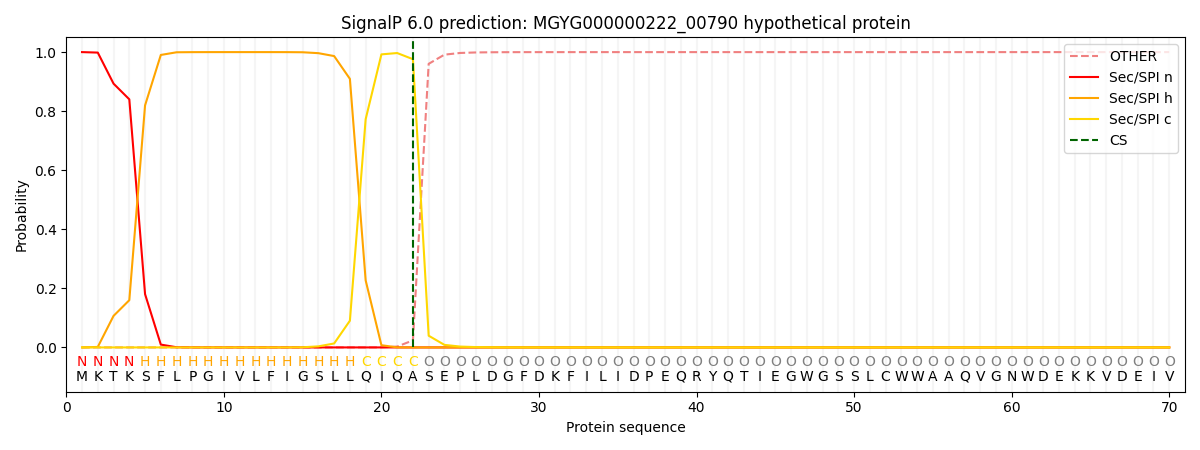
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000358 | 0.998922 | 0.000216 | 0.000164 | 0.000158 | 0.000148 |
