You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000000222_01755
You are here: Home > Sequence: MGYG000000222_01755
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Parabacteroides sp003473295 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Tannerellaceae; Parabacteroides; Parabacteroides sp003473295 | |||||||||||
| CAZyme ID | MGYG000000222_01755 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | Muramidase-2 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 125144; End: 125647 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1705 | FlgJ | 4.23e-25 | 16 | 163 | 60 | 181 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| PRK05684 | flgJ | 1.67e-20 | 4 | 163 | 153 | 303 | flagellar assembly peptidoglycan hydrolase FlgJ. |
| smart00047 | LYZ2 | 2.42e-16 | 4 | 163 | 9 | 138 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
| PRK12709 | flgJ | 3.01e-16 | 23 | 163 | 197 | 318 | flagellar rod assembly protein/muramidase FlgJ; Provisional |
| pfam01832 | Glucosaminidase | 4.63e-15 | 18 | 116 | 12 | 88 | Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase. This family includes Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase EC:3.2.1.96. As well as the flageller protein J that has been shown to hydrolyze peptidoglycan. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM53324.1 | 1.08e-68 | 1 | 166 | 1 | 167 |
| BAR50235.1 | 1.40e-53 | 5 | 166 | 9 | 172 |
| QUB71804.1 | 3.52e-53 | 4 | 163 | 7 | 169 |
| BAR52236.1 | 1.73e-52 | 5 | 166 | 34 | 197 |
| VEI52671.1 | 1.06e-27 | 1 | 162 | 1 | 164 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5DN5_A | 1.01e-11 | 1 | 162 | 1 | 146 | Structureof a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_B Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_C Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 5DN4_A | 1.31e-11 | 1 | 162 | 1 | 146 | Structureof the glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 3VWO_A | 3.89e-11 | 4 | 157 | 2 | 142 | Crystalstructure of peptidoglycan hydrolase mutant from Sphingomonas sp. A1 [Sphingomonas sp. A1] |
| 2ZYC_A | 4.55e-11 | 4 | 157 | 3 | 143 | ChainA, Peptidoglycan hydrolase FlgJ [Sphingomonas sp. A1] |
| 3K3T_A | 3.30e-10 | 4 | 157 | 3 | 143 | E185Amutant of peptidoglycan hydrolase from Sphingomonas sp. A1 [Sphingomonas sp. A1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39046 | 2.07e-12 | 3 | 157 | 66 | 204 | Muramidase-2 OS=Enterococcus hirae (strain ATCC 9790 / DSM 20160 / JCM 8729 / LMG 6399 / NBRC 3181 / NCIMB 6459 / NCDO 1258 / NCTC 12367 / WDCM 00089 / R) OX=768486 GN=EHR_05900 PE=1 SV=1 |
| Q9I4P4 | 2.83e-11 | 2 | 162 | 238 | 386 | Peptidoglycan hydrolase FlgJ OS=Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) OX=208964 GN=flgJ PE=3 SV=1 |
| P58231 | 3.04e-11 | 24 | 162 | 174 | 292 | Peptidoglycan hydrolase FlgJ OS=Escherichia coli O157:H7 OX=83334 GN=flgJ PE=3 SV=1 |
| P75942 | 3.04e-11 | 24 | 162 | 174 | 292 | Peptidoglycan hydrolase FlgJ OS=Escherichia coli (strain K12) OX=83333 GN=flgJ PE=3 SV=1 |
| P15931 | 5.16e-10 | 4 | 162 | 153 | 295 | Peptidoglycan hydrolase FlgJ OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=flgJ PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
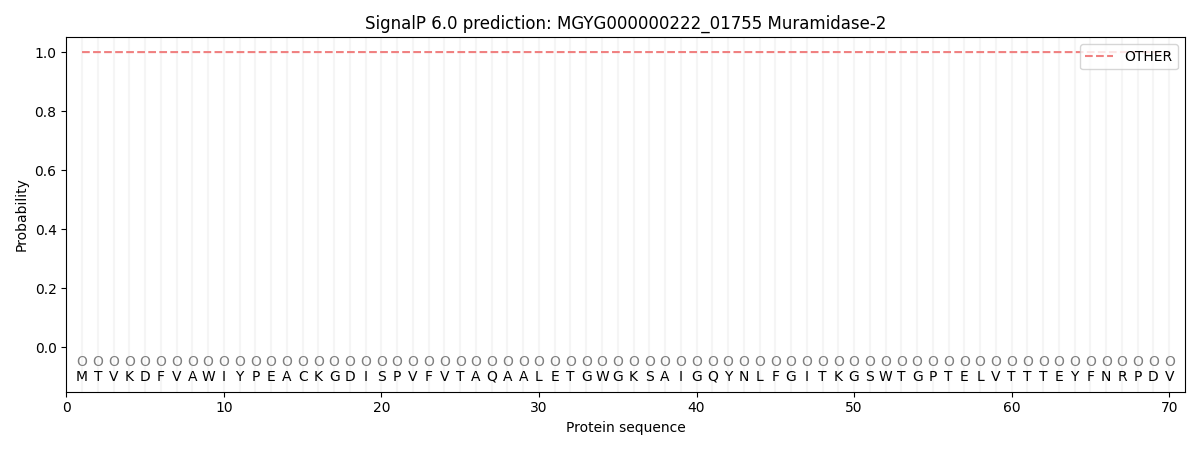
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000057 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
